The implications of rumen microorganism metabolism on environmental research
Introduction
The rumen is the first part of the complex four-component digestive system of the herbivores known as ruminants [1]. Over time, anaerobic microorganisms have grown in symbiosis with the ruminant inside the rumen compartment, breaking down recalcitrant plant polymers for the ruminant. This symbiosis allows the ruminant to absorb the resulting nutrients, while the herbivore provides the microorganism shelter and a diet [1]. Since these rumen microorganism enzymes can degrade recalcitrant plant polymers [2], there is interest in utilizing these microorganisms to solve environmental problems.
The Rumen
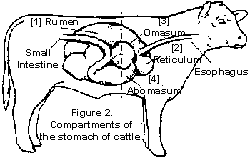
The rumen is a largely anaerobic environment that is maintained at a relatively constant temperature of 39°C [4]. Thus, fermentation is the key energy producing pathway. The physiology of the rumen depends largely on the ruminant’s diet: if the diet is deficient in fibre, fermentation increases, and rumen pH decreases [5]. As a result, the more acid-tolerant microorganisms such as Streptococcus bovis are more prevalent. S. bovis produces more lactic acid the faster it grows, thus there is positive feedback and the rumen becomes even more acidic, and acidosis can occur, which can be fatal to the ruminant [4].
Microbial Processes and Biological Interactions
Plant polymer degradation
A major component of the majority of plant matter is lignocellulose, a plant polymer. Though specific microorganisms are important in plant polymer degradation, such as fungi extracellularly degrading cellulose [8], it is the cumulative effect of various microorganisms that result in effective digestion [7]. For example, crossfeeding occurs between Selenomonas ruminantium and Bacteroides succiogenes. When cultured in vitro on a cellulose-containing plate, though S. ruminantium cannot degrade cellulose, it still grows on the plate, suggesting that B. succiogenes breaks the cellulose down into substrates that S. ruminantium can use [4]. This relationship is one of many commensal or mutualistic interactions that occur in the rumen.
Fermentation
Anaerobic protozoa ferment the plant polymers intracellularly into their respective monomers [4]. These are further broken down to volatile fatty acids such as acetate and butyrate [9], carbon dioxide (CO2), and hydrogen gas (H2), the latter two being used by methanogens to produce methane [4]. Volatile fatty acids are used by the ruminant as a source of energy [4].
Methanogenesis
A key process in the bovine rumen is methanogenesis by methanogens, the only known rumen methanogens being archaea [10]. Methanogens are unique in that methane production is coupled with ATP synthesis [10]. By using the fermentation end products CO2 as the carbon source and H2 as the electron source, as well as methyl-containing compounds and acetate [7], methanogens can produce methane.
Other pathways
Other pathways that microorganisms in the rumen facilitate are acetogenesis, sulfate reduction, nitrate reduction, and fumarate and propionate production [7].
Key Microorganisms
Bacteria
Bacteria are abundant in the rumen, with numerous varieties performing all of the important biochemical transformations in the rumen [5]. Ruminococcus albus, Ruminococcus flaveaciens, and Fibrobacter succinogenes represent the most active cellulolytic bacteria in the rumen [5]. Due to their ubiquity in the rumen and diverse functions, they are crucial in digestion of plant matter.
Protozoa
The protozoa in the rumen are predominantly ciliated protozoa, though there are also flagellated protozoa [7]. These are responsible for the primary breakdown of organic matter, engulfing it and converting it into volatile fatty acids, CO2, and H2. Protozoa consume bacteria and take up starch granules, affecting fermentation rate and acidity [5]. Interestingly, protozoa are not necessary for the survival of the ruminant, as defaunation (removal of the protozoa) is frequently used to study the rumen microbiota [9].
Fungi
Fungi in the rumen have also been shown to play a role in primary lignocellulose degradation, due to the presence of extracellular cellulases and xylanases [8]. Furthermore, unlike bacteria, archaea and protozoa, fungi have rhizoids that allow them to access areas that the other microorganisms cannot reach [11], adding to their functionality as lignocellulose degraders.
Archaea
The predominant methanogenic archaea present in the rumen are Methanobrevibacter spp., making up approximately two-thirds of the archaea population, while the other third is composed of mostly Methanomicrobium [7] and a group of archaea with unknown physiology termed rumen cluster C [12]. The archaea’s main role in the rumen is methanogenesis, the substrates of which are provided by fermentation by the bacteria and protozoa.
Environmental Impact and Current Research
Greenhouse gas reduction
Due to the large size of the dairy sector, the methane and CO2 contribute significantly to global greenhouse gas levels. Livestock emissions are estimated to contribute 9 percent of global anthropogenic carbon dioxide emissions and 35-40 percent global anthropogenic methane emissions [13]. Thus, there is ongoing research on how to reduce greenhouse gas emissions from cattle. One possible method is to change the diet of cattle: replacing alfalfa silage with corn silage was shown to reduce methane production for example [14]. However, it has been found that, methane expulsion is reduced, due to farming regulations, overall it will take over forty years for the crop change to pay off [3].
Metagenomics
Although some aspects of the general mechanisms of plant polymer breakdown are known, much is still uncertain, because currently many microorganisms native to the rumen cannot be cultured outside of the rumen [2]. Through the relatively recent development of metagenomics, many cellulose-active enzymes and their functions have been discovered [2]. For example, it was found that cellulosome complexes initially attack side chains of the plant polysaccharides, rather than the more recalcitrant main chains [15].
The search for biofuels
An extension of metagenome research is the search for an effective method for the breakdown of lignocellulic biomass. Rumen microorganisms are very efficient at degrading plant matter, and thus are a current area of research for producing biofuel from organic matter to act as renewable alternative energy sources [2]. For example, rumen microorganisms such as Clostridium have been shown to be able to degrade lignocellulose, such as in napier grass [16], switchgrass and miscanthus [2]. If successful, biofuels degradation by rumen microorganisms on an industrial scale could act as a very clean and efficient alternative to fossil fuels.
References
1. Hobson, P., Stewart, C. “The Rumen Microbial Ecosystem.” 1997, 10.1007/978-94-009-1453-7
2. Hess, M., Sczyrba, A., Egan, R., Kim, T., Chokhawala, H., Schroth, G., Luo, S., Clark, D., Chen, F., Zhang, T., Mackie, R., Pennacchio, L., Tringe, S., Visel, A., Woyke, T., Wang, Z., Rubin, E. “Metagenomic Discovery of Biomass-Degrading Genes and Genomes from Cow Rumen.” Science, 2011, DOI: 10.1126/science.1200387.
3. Hall, J., Silver, S. “Nutrition and Feeding of the Cow-Calf Herd: Digestive System of the Cow.” 2009, http://pubs.ext.vt.edu/400/400-010/400-010.html#TOC
4. Russell, J., Hespell, R. “Microbial Rumen Fermentation.” J. Dairy Sci., 1981, DOI: 10.3168/jds.S0022-0302(81)82694-X
5. Russell, J., Rychlik, J. “Factors That Alter Rumen Microbial Ecology.” Science, 2011, DOI: 10.1126/science.1058830.
6. Kowsari, M. “Pictures for Lignocellulose Structure.” 2012, http://biofuel.webgarden.com/sections/blog/pictures-for-lignocellulose
7. Morgavi, D., Forano, E., Martin, C., Newbold, C. “Microbial ecosystem and methanogenesis in ruminants.” Animal, 2010, DOI: 10.1017/S1751731110000546.
8. Bhat, M., Bhat, S. “Cellulose degrading enzymes and their potential industrial applications." Biotechnol. Adv., 1997, DOI: 10.1016/S0734-9750(97)00006-2.
9. Williams, A., Coleman, G. “The Rumen Protozoa.” 1992, 10.1007/978-1-4612-2776-2
10. Sirohi, S., Pandey, N., Singh, B., Puniya, K. “Rumen methanogens: a review.” Indian J. Microbiol., 2010, DOI: 10.1007/s12088-010-0061-6
11. Bauchop, T. “The anaerobic fungi in rumen fibre digestion.” Agriculture and Environment, 1981, DOI: 10.1016/0304-1131(81)90021-7.
12. Jeyanathan, J., Kirs, M., Ronimus, R., Hoskin, S., Janssen, P. “Methanogen community structure in the rumens of farmed sheep, cattle and red deer fed different diets.” FEMS Microbiol. Ecol., 2011, DOI: 10.1111/j.1574-6941.2011.01056.x.
13. Van Middelaar, C., Berentsen, P., Dijkstra, J., De Boer, I. “Evaluation of a feeding strategy to reduce greenhouse gas emissions from dairy farming: The level of analysis matters.” Agricultural Systems, 2013, doi: 10.1016/j.agsy.2013.05.009.
14. Hassanat, F, Gervais, R, Julien, C, Massé, DI, Lettat, A, Chouinard, PY, Petit, HV, Benchaar, C. 2013. “Replacing alfalfa silage with corn silage in dairy cow diets: Effects on enteric methane production, ruminal fermentation, digestion, N balance, and milk production.” J. Dairy Sci., 2013, doi: http://dx.doi.org/10.3168/jds.2012-6480.
15. Ho, C., Chang, J., Lin, J., Chin, T., Mathew, G., Huang, C. “Establishment of functional rumen bacterial consortia (FRBC) for simultaneous biohydrogen and bioethanol production from lignocellulose.” Int J Hydrogen Energy, 2011, DOI: 10.1016/j.ijhydene.2011.06.125.
16. Food and Agriculture Organization of the United Nations (FAO). “Livestock’s Long Shadow: Environmental Issues and Options”. 2006, http://www.fao.org/docrep/010/a0701e/a0701e00.htm