Thermoplasma
A Microbial Biorealm page on the genus Thermoplasma
Classification
Higher order taxa:
Archaea; Euryarchaeota; Thermoplasmata; Thermoplasmatales; Thermoplasmataceae
Species:
Thermoplasma acidophilum, T. volcanium
Description and Significance
Thermoplasma was first isolated in the 1960s, in a coal pile in Indiana. Thermoplasma is a thermophile, meaning that it grows best in hot environments, usually between 55 and 60 degrees Celsius. This genus is most famous for its acidophilia, preferring pH range of 0.5-4. Thermoplasma cells lyse at a neutral pH.
Genome Structure
In 2001, the genome strucutres of both Thermoplasma acidophilum DSM 1728 and Thermoplasma volcanium GSS1 were sequenced. Until the genome was sequenced, Thermoplasma was believed to belong to the Eukaryotes. T. acidophilum contains 1,564,905 base pairs; it is one of the smallest genomes ever sequenced. Despite its size, there are 68 proteins that do not exist in any other archaeal genome. This genome lacks some characteristics as well. For example, although Thermoplasma have flagella, no chemotaxis protein has been found. In addition, Thermoplasma genes that reduce sulfur resemble those of bacteria instead of other archaea.
Cell Structure and Metabolism
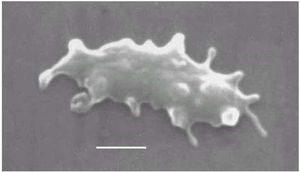
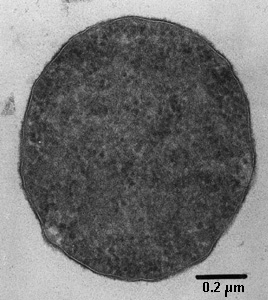
Like all archaea, Thermoplasma are unicellular. Despite living in acidic environments, Thermoplasma lack cell walls. They are contained only by plasma membranes which consists of diether and tetraether lipids. In addition, Thermoplasma are not nucleated. The shape of these cells can be filamentous or coccoid, or they can take the form of a disc or club. Thermoplasma are flagellated. Thermoplasma show an irregular form when placed in water, and become spheres at room temperature.
Thermoplasma are heterotrophic organisms. They are capable of obtaining energy in anaerobic environments via sulphur respiration. Thermoplasma are also scavengers, eating organisms that cannot survive in a highly acidic environment.
Ecology

Thermoplasma are found in self-heating coal waste piles, although this is not a natural habitat. They are typically found in hot springs, as well as solfataric fields. Thermoplasma acidophilum can be employed as a model organism for scientific research, partially because this species has a strong evolutionary relationship to eukaryotes.
References
DeLong, Edward F. "Extreme Genomes." Genome Biology. 2000, 1(6):1029.1–1029.3.
Mayr, Jutta, Huei-Rong Wang, Petra Nederlof, and Wolfgang Baumeister."The Import Pathway of Human and Thermoplasma 20S Proteasomes into HeLa Cell Nuclei Is Different from That of Classical NLS-Bearing Proteins." Biol. Chem., Vol. 380, pp. 1183 – 1192, October 1999.
Thermoplasma Genome Project Pages. 12 February 2002. Accessed 4 August 2005.