User:S4342592
Nur' Amirah Mohd Kassim Bench D 09082016
Classification
Higher Order Taxa
- Kingdom: Bacteria
- Domain: Bacteria
- Phylum: Bacteroidetes
- Class: Bacteroidia
- Order: Bacteroidales
- Family: Porphyromonadaceae
- Genus: Porphyromonas
Species
Porphyromonas gingivalis strain 2561 = ATCC 33277= CCUG 25893 = CCUG 25928 = CIP 103683 = DSM 20709 = JCM 12257 = NCTC 11834.

Description and Significance
Porphyromonas gingivalis is a gram negative, non motile, rod shaped bacteria that is pathogenic, with no functional role. It is commonly found in the oral cavity. On blood agar plate, it forms black pigmented colonies due to the build-up of hemin that helps to transport iron [1]. P. gingivalis is the main source of periodontal disease and is also said to be associated with cardiovascular disease and diabetes amongst other inflammatory disease thus it is important to study this bacterium [2].
Genome Structure
In 2003, the genome of P. gingivalis was described to have 2,343,479bp with an average guanine cytosine nucleotide content of 48.3%. It has 4 ribosomal operons, 2 RNA structural genes and 53 tRNA gene. There was a total of 1990 open reading frames that could be identified in the genome. 21 areas of the genome were discovered to display an atypical nucleotide composition. The areas range in size from 11 to 68kb and has a guanine cytosine nucleotide content of between 29.4% to 61.6%. 463 genes were also seen in the genome [3][4]
Cell Structure and Metabolism
Cell Wall
To get established in the oral cavity, microorganism have to adhere to the teeth surface. Adhesins are molecules found in the capsule of the cell wall that allows the adhesion. The capsule is therefore responsible in the initial adhesion to the human periodontal pocket epithelial cells, where it helps facilitate attachment. The outer membrane that contains the fimbriae and LPS amongst other structures, is an asymmetrical bilayer that plays a role as a selective barrier that offers protection and allow movement of various substrate through the outer membrane porin proteins. Fimbriae are thin, proteinaceous surface appendages that protrudes from the outer membrane of a bacterial cell. Its function is essential for invasion of host cells, adherence to oral substrates as well as contribution to the progression of periodontal inflammatory reactions. Lipopolysaccharide (LPS) is also an important component of the bacterial outer membrane. It helps to facilitate bacterial survival and maintain the cellular and structural integrity as well as controlling the entry of hydrophobic molecules and toxic chemicals. [5].
Biofilm Formation
Dental plaque develops on tooth surfaces and in certain circumstances, it progresses to a more pathogenic condition such as periodontal diseases. The process of plaque build-up is characterised by binding interactions of various oral bacteria. Early, primary commensal colonisers such as oral streptococci attach to the enamel pellicle, covering the tooth surface and providing attachment sites for subsequent colonisation by other potential pathogenic bacteria such as P. gingivalis. Evidently, together with Streptococcus gordonii, P. gingivalis is able to colonise efficiently, making it an important process in biofilm formation [6].
Metabolic Functions
Genome analysis suggest various metabolic pathways that P. gingivalis is capable of. There are also some metabolic pathways that P. gingivalis perform poorly or not able to perform. P. gingivalis has a limited capability for organic nutrient metabolism and poor glucose utilisation. In addition, carbohydrates do not support the growth of the bacteria. However, P. gingivalis contain ORF for all enzymes that is necessary for glycolytic pathway such as glucose/galactose transporters and glucose kinase. Pentose phosphate pathway was identified for anaerobic growth. Amino acid is also available for energy production. Other pathways that P. gingivalis is capable of performing include lysine catabolic pathway and iron acquisition [7].
Ecology
P. gingivalis is an anaerobic rod shaped microorganism that is mainly found in the subgingival plaque of the periodontal in the oral cavity. Other potential environment where the pathogenic bacteria can be found include the upper gastrointestinal tract, the colon and the respiratory tract [3]. It has also been found in women with bacterial vaginosis [8]. P. gingivalis has the ability to actively internalise within the epithelial cells by mechanism similar to various enteric pathogens, other than being capable of penetrating gingival tissues. The epithelial cells comprises of interactive interface that generate and transmit signals between bacteria and the adjacent immune cells in the periodontal tissues. Many bacterial species initiate infections at epithelial cell interfaces to internalise within the epithelial cells. [9]
Pathology
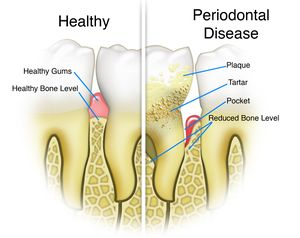
P. gingivalis is responsible for causing periodontal disease. Periodontal disease is characterized by the destruction of tooth supporting tissues due to complex polymicrobial infection. The disease progresses from acute inflammation of the gingival tissue to formation of tooth pockets which eventually leads to the loss of teeth. Symptoms of periodontal disease include swollen or bleeding gums, sensitive teeth and receding gums [5] Other than causing periodontal disease, the microorganism has also been associated with rheumatoid arthritis, which is an autoimmune disease. This is due to the correlation that anti-P. gingivalis antibodies were more frequently observed in rheumatoid arthritis patients compared to healthy controls. There was also a significant amount of bacterial DNA detected in the synovial fluid that indicates the transfer of bacterial DNA from the infection site to the joints. [10]
Application to Biotechnology
Given the emerging evidence that periodontal infections and systemic conditions have been closely associated with one another, the search for a method to detect the presence of P. gingivalis is therefore necessary. Characterised antibodies have been raised against purified P. gingivalis HmuY protein and elected epitopes of the HmuY molecule. The results obtain from the study suggest that P. gingivalis HmuY protein may be used as an antigen for antibodies to target this bacteria. Therefore, purified HmuY protein will be use to suggest levels of anti-P. gingivalis antibodies in the patients' sera with chronic periodontitis [11]. In another study, various plant derived compounds such as quercetin and resveratol has shown inhibitory effects on the activity of P. gingivalis fimbriae, however, further studies needs to be conducted in order to finally produce an effective inhibitor against P. gingivalis [12].
Current Research
Current research have been investigating the link between cardiovascular disease (CVD) and P. gingivalis. There is evidence of epidemiological association between oral infection and CVD thus studies have indicated the plausible link between P. gingivalis infection and atherosclerosis. Intervention studies have also demonstrated that periodontal therapy can reduce the systemic markers of inflammation and vascular health. Though further and more in depth studies are needed, it is recommended that patients with moderate to severe cases of periodontitis be informed of the the possible risk of CVD [2].
References
13. Comparison of Healthy Gum vs. Periodontal Disease Gum
This page is written by Nur' Amirah Mohd Kassim for the MICR3004 course, Semester 2, 2016
