Ensifer adhaerens: Difference between revisions
No edit summary |
|||
| Line 38: | Line 38: | ||
E. adhaerens is capable of binding to a range of Gram-Positive and Gram-Negative bacterial cells; however, the host range of E. adhaerens appears to be limited to primarily Gram-Positive bacteria, which it is capable of lysing [2.3]. Attachment to prey bacteria appears to involve appendages referred to as “Bars” by L.E. Casida in the initial publications detailing the predatory process [3]; the purpose of this appendage is, as of yet, unknown. It is possible that bars function in attachment or promotion of nutrient diffusion to Ensifer cells, however neither of these hypotheses have been assessed. Much of what is known about the predatory habits of Ensifer adhaerens was worked out in the early 1980’s, and few molecular details of the process are known. Remaining areas of study regarding Ensifer predation include what genetic potential is required to act as a bacterial predator, as well as the molecular details of the predatory process. With the recent availability of the Casida strain genome, this area can be studied in greater detail with modern molecular genetics [6]. Potential areas to explore this process in greater detail include transcriptomics and forward genetics screens to determine patterns of gene expression during predation and essential predation genes, respectively. | E. adhaerens is capable of binding to a range of Gram-Positive and Gram-Negative bacterial cells; however, the host range of E. adhaerens appears to be limited to primarily Gram-Positive bacteria, which it is capable of lysing [2.3]. Attachment to prey bacteria appears to involve appendages referred to as “Bars” by L.E. Casida in the initial publications detailing the predatory process [3]; the purpose of this appendage is, as of yet, unknown. It is possible that bars function in attachment or promotion of nutrient diffusion to Ensifer cells, however neither of these hypotheses have been assessed. Much of what is known about the predatory habits of Ensifer adhaerens was worked out in the early 1980’s, and few molecular details of the process are known. Remaining areas of study regarding Ensifer predation include what genetic potential is required to act as a bacterial predator, as well as the molecular details of the predatory process. With the recent availability of the Casida strain genome, this area can be studied in greater detail with modern molecular genetics [6]. Potential areas to explore this process in greater detail include transcriptomics and forward genetics screens to determine patterns of gene expression during predation and essential predation genes, respectively. | ||
==References== | ==References== | ||
Revision as of 18:30, 30 April 2020
Classification
Bacteria > Proteobacteria > Alphaproteobacteria > Rhizobiales > Rhizobiacea
Species
|
NCBI: Taxonomy |
Ensifer adhaerens
Description and Significance
E. adhaerens is a soil-dwelling bacteria belonging to the Alphaproteobacteria in the Rhizobiales family [1,2,3]. E. adhaerens survives in the soil as a facultative predator of soil bacteria, and was initially identified in screens to determine a microbial factor for Micrococcus luteus'’ inability to grow in soil [2,3,4].Multiple facets of E. adhaerens biology are being exploited for biotechnology applications, including the ability of E. adhaerens to engage in plant transformation when supplied with the Agrobacterium tumefaciens Ti plasmid and E. adhaerens’' ability to degrade volatile organic compounds [8,9,10].
Ensifer-mediated transformation is of interest due to patent circumvention of Agrobacterium-mediated transformation methods, and potential increases in transformation efficiency [10]. This approach is similar in principle to Agrobacterium-mediated transformation, in that the Transfer DNA region of an Agrobacterium Ti plasmid is engineered to contain a gene of interest, and this plasmid is moved into E. adhaerens [10]. E. adhaerens then expresses the machinery necessary for the transfer of the T-DNA segment into the plant cell, which then recombines into the plant chromosome [10]. Transformation efficiencies comparable to Agrobacterium have been obtained in some strains of Japonica rice, for example, though Ensifer-mediated transformation efficiency lags Agrobacterium for other strains [10]. Recent transcriptomic experiments on transformation-capable E. adhaerens strains may be used to guide further optimization of Ensifer-mediated transformation protocols [11].
Another potential application for E. adhaerens is in bioremediation [9,10]. E. adhaerens isolates have been shown to break down complex organic pollutants, and environmental isolates are promising for their potential to serve as bioremediation tools [9,10]. The TMX-23 strain of E. adhaerens, for example is capable of breaking down the pesticide TMIA [10]. In addition, the TMX-23 strain also produces secondary metabolites that are beneficial to plant growth and germination [10]. An appealing scenario for the use of this strain would be to seed the rhizosphere surrounding crop plants with this microbe concordant with the use of the TMIA pesticide; this would prevent accumulation of pesticide in the soil and promote the growth of plants in the soil [10].
Genome Structure
The full-genome sequences of multiple isolates of Ensifer adhaerens have been published [6]. The E. adhaerens isolate Casida A, the first strain of E. adhaerens to be characterized, sequence reveals five replicons totaling 7.2 Mbp and 6641 open reading frames [6]. The 16s rRNA gene places E. adhaerens near the Sinorhizobia phylogenetically, and there is some taxonomic debate as to whether E. adhaerens should be placed within the Sinorhizobia [1,7]. Regardless, the E. adhaerens literature typically resorts to the Ensifer adhaerens name and not Sinorhizobium adhaerens.
Cell Structure, Metabolism and Life Cycle
E. adhaerens is a terrestrial, rod-shaped Alphaproteobacteria that belongs to the Rhizobiales family [1,2,3]. E. adhaerens survives in the soil as a facultative predator of soil bacteria, and was initially identified in screens to determine a microbial factor for Micrococcus luteus’ inability to grow in soil [2,3,4]. E. adhaerens is capable of growth on glucose as a sole substrate in buffered media containing salts, indicating that the predatory behavior of E. adhaerens is not obligatory and that E. adhaerens is biosynthetically competent [2].
As with several other members of the Alphaproteobactiera, Ensifer ahdaerens undergoes a polar budding cell division process [2]. This process is common to Agrobacterium, Rhodopseudonas, and Sinorhizobium [5]. While the genetic and biochemical factors involved in E. adhaerens cell cycle have not been directly studied, the Rhizobial cell division process is a current topic of research and mirrors the effort to characterize the Alphaproteobacteria Caulobacter crescentus cell cycle.
Ecology and Pathogenesis
E. adhaerens is a resident of the rhizosphere, the layer of soil influenced by the roots of plants. E. adhaerens is thought to survive partially as a bacterial predator, though E. adhaerens is capable of surviving in the absence of host cells [2]. Additionally, as evidenced by the close evolutionary relationship of E. adhaerens to the Sinonrhizobia, some isolates have been recovered that are capable of nitrogen fixation in root nodules [10].
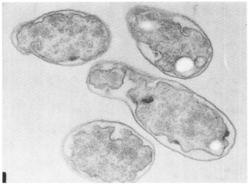
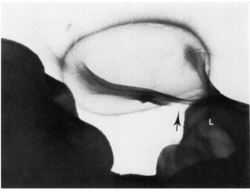
E. adhaerens is capable of binding to a range of Gram-Positive and Gram-Negative bacterial cells; however, the host range of E. adhaerens appears to be limited to primarily Gram-Positive bacteria, which it is capable of lysing [2.3]. Attachment to prey bacteria appears to involve appendages referred to as “Bars” by L.E. Casida in the initial publications detailing the predatory process [3]; the purpose of this appendage is, as of yet, unknown. It is possible that bars function in attachment or promotion of nutrient diffusion to Ensifer cells, however neither of these hypotheses have been assessed. Much of what is known about the predatory habits of Ensifer adhaerens was worked out in the early 1980’s, and few molecular details of the process are known. Remaining areas of study regarding Ensifer predation include what genetic potential is required to act as a bacterial predator, as well as the molecular details of the predatory process. With the recent availability of the Casida strain genome, this area can be studied in greater detail with modern molecular genetics [6]. Potential areas to explore this process in greater detail include transcriptomics and forward genetics screens to determine patterns of gene expression during predation and essential predation genes, respectively.
References
- Rogel, MA; et al. Nitrogen-Fixing Nodules with Ensifer adhaerens Harboring Rhizobium tropici Symbiotic Plasmids. 2001. Applied and Environmental Microbiology 67(7):3264-3268
- Casdia, LE. Bacterial Predators of Micrococcus luteus in Soil. 1980. Applied and Environmental Microbiology 39(5): 1035-1041.
- Casida, L.E. Jr. Ensifer adhaerens gen. nov., sp. nov.: a Bacterial Predator of Bacteria in Soil. 1982. International Journal of Systematic Bacteriology 32(3):339-345.
- Germida, JJ; Casida, LE. 1983. Ensifer adhaerens Predatory Activity Against Other Bacteria in Soil, as Monitored by Indirect Phage Analysis. Applied and Environmental Microbiology 45(4):1380-1388.
- Brown, PJB; et al. Polar Growth in the Alphaproteobacterial order Rhizobiales. 2012. Proceedings of the National Academy of Science 109(5): 1697-1701.
- Williams, LE; et al. Complete Genome Sequence of the Predatory Bacterium Ensifer Adhaerens. 2017. Microbiology Resource Announcements
- Young, JM. The Genus Name Ensifer Casida 1982 Takes Priority Over Sinorhizobium Chen Et Al. 1988, and Sinorhizobium morelense Wang Et Al. 2002 is a Later Synonym of Ensifer adhaerens Casida 1982. Is the Combination “Sinorhizobium adhaerens” (Casida 1982) Willems Et Al. 2003 Legitimate? Request for an Opinion. 2003. Int J Syst Evol Microbiol 53(6):2107-2110.
- Xu, L; et al. Characterization of the Biosorption and Biodegradation Properties of Ensifer adhaerens: A Potential Agent to Remove Polychlorinated Biphenyls from Contaminated Water. 2016. Journal of Hazardous Materials 302: 314-322.
- Zhou, G; et al. Biodegradation of the neonicotinoid insecticide thiamethoxam by the nitrogen-fixing and plant-growth promoting rhizobacterium Ensifer adhaerens strain TMX-23. 2013. Applied Microbial and Cell Physiology 97:4065-4074.
- Zuniga-Soto, E; et al. Ensifer-mediated Transformation: An Efficient non-Agrobacterium Protocol for the Genetic Modification of Rice. 2015. SpringerPlus 4:600.
- Zuniga-Soto, E; et al. Insights into the Transcriptomic Response of the Plant Engineering Bacterium Ensifer adhaerens OV14 During Transformation. 2019. Science Reports 9(1):10344.
Author
Page authored by Joseph Stembel, student of Prof. Jay Lennon at IndianaUniversity.