Glycylcycline Antibiotics: Difference between revisions
No edit summary |
No edit summary |
||
| (26 intermediate revisions by one other user not shown) | |||
| Line 1: | Line 1: | ||
{{Curated}} | |||
By: Kristina Buschur | |||
==Introduction== | ==Introduction== | ||
<br>The ability of bacteria to quickly develop resistance to commonly used antibiotics is a huge hurdle in the path of disease treatment. Because of this, there is an ever-present need to develop new antibiotics that use novel mechanisms to overcome multidrug-resistance and prevent microbial growth. Since tetracycline has been in wide use since the mid-1900s for treatment of many human and animal infections and as growth promoters in agriculture, many bacteria have since developed sophisticated mechanisms to prevent the harmful effects of the antibiotic (Greer 2006). Specifically, tetracycline-speicific efflux pumps and ribosomal protection proteins are commonly employed by bacteria. The glycylcycline class of antibiotics is a powerful tool that has been recently developed to combat this problem. Developed by Lederle Laboratories (now American Home Products) in the early 1990s and derived from tetracycline, glycylcyclines have added substituents that interfere with the mechanisms bacteria employ to resist tetracycline, including both the efflux pumps and ribosomal protection proteins (Chopra 2002). <br><br> | |||
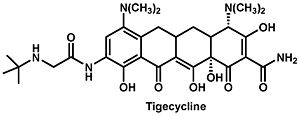
[[Image:tigecycline.jpeg|thumb|300px|right|The structure of tigecycline.]] | [[Image:tigecycline.jpeg|thumb|300px|right|Figure 1. The structure of tigecycline (Olson 2006).]] | ||
Currently tigecycline (previously GAR-936) is the only antibiotic of the glycylcycline class in clinical use. Tigecycline was approved by the Food and Drug Administration in 2005, and is particularly useful in the treatment of multi-drug resistant infections, which are especially hard to | Currently tigecycline (previously GAR-936) is the only antibiotic of the glycylcycline class in clinical use (Figure 1). Tigecycline was approved by the Food and Drug Administration in 2005, and is particularly useful in the treatment of multi-drug resistant infections, which are especially hard to eradicate. Penicillin-resistant ''Streptococcus pneumoniae'', methicillin-resistant ''Staphylococcus aureus'' and ''Staphylococcus epidermidis'', and vancomycin-resistant ''Enterococcus'' species are several examples of species that have developed an antibiotic resistance but are still affected by tigecycline (Greer 2006). In addition to these, there is a wide variety of organisms, including both gram-positive and gram-negative bacteria, that are sensitive to tigecycline. It does not, however, seem to have any effect on ''Proteus'', ''Providencia'', or ''Pseudomonas'' species. Even still, tigecycline shows particular success against some anaerobic organisms including ''Bacteriodes fragilis'', ''Peptostreptococcus'' species, and ''Propionibacterium acnes'' which for the most part are able to evade tetracycline. <br><br> | ||
The tigecycline antibiotic is structurally very similar to minocycline and similarly binds to the bacterial 30S ribosome unit. The manner in which the molecule binds prevents amino-acyl tRNAs from binding to the A site of the ribosome and subsequently prevents peptide formation and bacterial growth. The main difference between tigecycline and minocycline is the addition of an N,N-dimethylglycylamido group which actually causes the molecule to bind to the ribosome up to five times more tightly which in turn decreases the probability that resistance will develop. In general, the glycylcycline class of antibiotics (the 9-glycinyltetracyclines) is characterized by a molecular structure containing a four-ring carbocyclic skeleton with a substitution of an N-alkylglycylamido group at the D-9 position (Bhattacharya 2009). Biochemical experiments have shown that the tigecycline molecule binds to the same site on 16S rRNA as tetracycline but in a different orientation and with greater affinity. This has been confirmed by experiments in which a decreased binding affinity was observed in strains of ''E. coli'' with known mutations in the A site of the rRNA (G966U or G1058C). <br><br> | |||
Tigecycline is administered to a patient intravenously with a dose of 50 mg every 12 hours after an initial 100 mg loading dose. As with nearly all drugs, | Tigecycline (marketed as Tygacil) is used for the treatment of complicated skin and skin structure infections and complicated intraabdominal infections caused by susceptible organisms. The drug is administered to a patient intravenously with a dose of 50 mg every 12 hours after an initial 100 mg loading dose (Greer 2006). This dosage is recommended for patients regardless of age, sex, or race. After being administered, it is found at a higher concentration in the gallbladder, lungs, and colon than in the serum. In the bone and synovial fluid, though, it is found in lower concentrations than in the serum. Its elimination half-life is about 36 hours, and it leaves the body mainly through the bile and feces, but partly through the urine as well. As with nearly all drugs, tigecycline has several known side-effects, including nausea, vomiting, and diarrhea, but these are relatively minor. Barring any severe complications, the treatment period usually ranges from five to fourteen days (Bhattacharya 2009). <br><br> | ||
| Line 11: | Line 14: | ||
==Comparison of Tetracycline and Tigecycline Ribosome Binding== | ==Comparison of Tetracycline and Tigecycline Ribosome Binding== | ||
<br> Bauer et al. have identified the binding sites of tetracycline and tegecycline through probing 70S E. coli ribosomes with dimethylsulphate (DMS) and | <br> Bauer ''et al.'' have identified the binding sites of tetracycline and tegecycline through probing 70S ''E. coli'' ribosomes with dimethylsulphate (DMS) and Fe<sup>2+</sup>-mediated cleavage. This method works by allowing Fe<sup>2+</sup> to replace Mg<sup>2+</sup> complexed with tetracyclines. When H<sub>2</sub>O<sub>2</sub> is added, the Fe<sup>2+</sup> produces transient, very reactive hydroxy radicals that can cleave RNA close to where tetracycline is bound. It follows then, that the resulting cleavage sites will also be the location at which tetracycline binds to the ribosome (Bauer 2004). Crystallography experiments have shown that there are six tetracycline binding sites on the 30S ribosome. <br><br> | ||
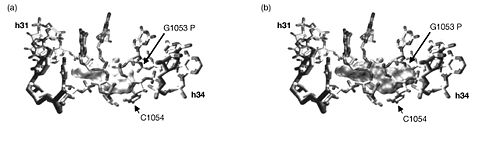
Three sites in the rRNA were found that were cleaved in the presence of tetracycline. Two of the three sites, those in h31 and h34, corresponded with tetracycline binding site-1. These were confirmed through a point mutation (G1058 to C) in h34 that produced tetracycline resistance. Additional mutation experiments showed that mutations in these areas also resulted in an eight-fold increase in resistance to tigecycline. The third rRNA cleavage site in h29 corresponds to tetracycline binding site-4, so the observation is confirmed by crystallography data. There is, however, no evidence of a magnesium ion complexed with tetracycline at tetracycline binding site-2. It follows, then, that the Fe<sup>2+</sup>-mediated cleavage technique would not reveal a tetracycline at this site. The other tetracycline binding sites have previously shown no evidence of being involved in antibiotic activity. <br><br> [[Image:16sbinding.jpeg|thumb|500px|right|Figure 2. Surface structures of tetracycline (a) and glycylcycline derivative DMG-DMDOT (b) bound to site-1. The bulky substituent on DMG-DMDOT overlaps with the phosphate backbone and prevents the molecule from binding in the same orientation that tetracycline does (Bauer 2004).]] | |||
With this knowledge, the authors conclude that since the three cleavage sites correspond to the physiologically important tetracycline binding sites and since the experiments produced identical results whether tetracycline or tigecycline was used, then tetracycline and tigecycline must share the same binding sites on 16S rRNA. Furthermore, tigecycline appears to bind much more tightly and at lower concentrations. There were, however, some differences in the Fe2+ cleavages intensities between tetracycline and tigecyclince. So although they do bind the ribosome in the same location, tigecycline likely binds in a different orientation than does tetracycline. The authors propose that the bulky t-butylglycylamido substituent at position 9 on tigecycline but | With this knowledge, the authors conclude that since the three cleavage sites correspond to the physiologically important tetracycline binding sites and since the experiments produced identical results whether tetracycline or tigecycline was used, then tetracycline and tigecycline must share the same binding sites on 16S rRNA. Furthermore, tigecycline appears to bind much more tightly and at lower concentrations. There were, however, some differences in the Fe2+ cleavages intensities between tetracycline and tigecyclince. So although they do bind the ribosome in the same location, tigecycline likely binds in a different orientation than does tetracycline. The authors propose that the bulky t-butylglycylamido substituent at position 9 on tigecycline but absent on tetracycline is responsible for this difference. To further investigate this, they compare the superimposed co-crystal structures of tetracycline and DMG-DMDOT (a glycylcycline derivative very similar to tigecycline) complexed with helices 31 and 34, which make up tetracycline binding site-1 (Figure 2). This comparison revealed that the bulky dimethylglycylamido substituent on DMG-DMOT would clash with the phosphate backbone of G1053, forcing it to bind the ribosome differently (Bauer 2004). Likewise, the t-butylglycylamido substituent on tigecycline is even bigger and would presumably change the orientation in which tigecycline binds in a similar, if not more dramatic, way. <br><br> | ||
<br> | <br> | ||
==Antibacterial Activity of Tigecycline== | ==Antibacterial Activity of Tigecycline== | ||
<br> | <br> | ||
[[Image:IVT.jpeg|thumb|400px|right|Figure 3. In vitro transcription/translation assays showing the effect of tigecycline (a), minocycline (b), and tetracycline (c) on GFP expression as a means to measure protein synthesis inhibition. IC<sub>50</sub> value anaylsis (d) shows that tigecycline has a higher percent inhibition of protein synthesis than minocycline and tetracycline over a wide range of concentrations. (Olson 2006).]] | |||
In 2006, Olson ''et al.'' reported using functional, biophysical, and molecular modeling experiments to determine the way in which tigecycline demonstrates antibacterial activity. To investigate the effect of tigecycline, in comparison with tetracycline and minocycline, on bacterial protein synthesis, they used in vitro transcription/translation (IVT) assays in which ''E. coli'' cell lysate was supplied with a GFP vector and the amount GFP produced was measured with a phosphorimager. The results clearly show a pattern in which the amount of GFP produced decreases quickly as the concentration of tigecycline, minocycline, or tetracycline is increased. This suggests that each of these molecules inhibits translation (Figure 3). Although all three antibiotics exhibit this trend, it is also important to note that tigecycline started inhibiting translation at the lowest concentration. At a tigecycline concentration of just 1.7 µM a significant inhibition of protein synthesis can be observed. This is in comparison to the much higher concentrations of minocycline (6.1 µM) and tetracycline (62.4 µM) that are required before any notable inhibition can be observed. These observations are summarized in Figure 3d, which presents a half maximal inhibitory concentration (IC<sub>50</sub>) anaylsis. The graph makes it clear that over a large range of concentrations, tigecycline has a much higher percent inhibition of protein synthesis than minocycline, which in turn is also more effective than tetracycline. This pattern also makes sense when it is considered that minocycline is a tetracyline derivative, and tigecycline is a minocycline derivative (Figure 4). <br><br> | |||
To further investigate the potency of these three drugs, competition studies very performed. They observed which of the antibiotics bound most effectively to both the 30S and the 70S subunits, and it was found that tigecycline bound to both the 30S and 70S subunits with 5-fold greater affinity than minocycline and 100-fold greater affinity than tetracycline (Olson 2006). Again, this trend is consistent with the structures of these molecules. <br><br> | |||
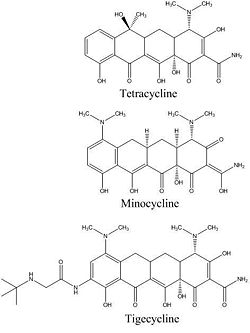
[[Image:antibioticcomparison.jpg|thumb|250px|left|Figure 4. Comparison of the structures of tetracycline, minocycline, and tigecycline.]] | |||
Finally, computational docking studies were performed to investigate the tigecycline-ribosome interaction. One particular position showed energetically favorable interactions that were consistent with all previous data. This position demonstrated favorable electrostatic, van der Waals, and hydrogen-bonding interactions with the 30S subunit that suggested it associated with the ribosome more closely than both minocycline and tetracycline. The overall orientation was very similar to that previously described for tetracycline, with the Mg ion interacting with the phosphate groups of G1197, C1054, and U1196. The most notable difference, though, was that the glycylcyclines interacted directly with other areas of the A site. These interactions included additonal hydrogen-bonding and van der Waals interactions that had never been seen before in that area (Olson 2006), observations that are consistent with the increased binding affinity of the glycylcyline class of antibiotics. <br><br> | |||
Combined, these observations further support the conclusion of Bauer ''et al.'' that tigecycline sterically inhibits the binding of aminoacyl-tRNA to the A site of the ribosome in a manner very similar to that of tetracycline. Furthermore, their data suggest that the way in which glycylcyclines are able to avoid TetM-mediated resistance could be the result of either their increased affinity for binding the ribosome or the unique orientation in which they bind the ribosome that might make TetM ineffective in protecting the ribosome. Perhaps, even, it is some combination of these two qualities that makes glycylcylines so effective.<br><br> | |||
<br><br> | |||
==Tigecycline Resistance Mechanisms== | |||
<br>Despite its current success in preventing growth of bacterial species that have already developed resistance to tetracycline, it is inevitable that bacteria will eventually mutate so that they are resistant to tigecycline as well. This is particularly a concern since tigecycline is so structurally similar to tetracycline, an antibiotic many bacteria have already developed successful defenses against. It is thought that resistance to tigecycline will develop even more quickly as a result of this (Chopra 2002). McAleese ''et al.'' have already found laboratory-derived mutants of ''Staphylococcus aureus'' that exhibit resistance to tigecycline and have traced that resistance to increased expression of the multidrug export protein ''mepRAB'' gene cluster. This cluster includes ''mepA'', a novel MATE family efflux pump. <br><br> | |||
While there are no known naturally occurring isolates of ''S. aureus'' that show resistance to tigecycline, two such mutants were found by serial passages through increasing concentrations of the antibiotic. One mutant was derived from the Mu3 strain (a clinical MRSA isolate with heterogeneous resistance to vancomycin) and the other from the N315 strain (a clinical MRSA isolate). The bacteria that resulted from this procedure had a very much reduced sensitivity to tigecycline. A susceptibility profile of the mutants was performed to determine if the sensitivity to other antibiotics had decreased as well, which might suggest a more general resistance mechanism was at work. They found that both of the mutants were more resistant to DMG-MINO and DMG-DMDOT, two other glycylcylcline antibiotics very similar in structure to tigecycline. Interestingly, the N315 mutant also showed increased resistance to EtBr and TPP (McAleese 2005). <br><br> | |||
A transcriptional profile analysis was undertaken to determine the origin of the decreased susceptibility observed in these mutants. ''S. aureus'' GeneChips were used to compare the DNA and RNA profiles of the Mu3 and N315 with their tigecycline-resistant mutants. They looked for changes in DNA content or gene expression that could explain the increased resistance. The found no changes in the DNA present in the mutants from the parent strains, suggesting that the mutants hadn't lost any genes that might be causing them to be susceptible to the antibiotic. The technique, though, was not sensitive enough to detect point mutations or small deletions, so it is still possible that these changes had occured. RNA was harvested from the parent and mutant strains during mid-exponential growth to compare their transcriptional profiles. Fifteen genes were found (though some of them are only hypothetical) whose transcription saw at least an eight-fold difference between the parent and the mutant (Figure 5). Several of these genes that are believed to code hypothetical proteins and other genes provided functions for which a connection to tigecycline resistance was not immediately apparent. Among these fifteen genes, though, was ''mepA'', an efflux pump-encoding gene. Additionally, ''tetM'', a tetracycline resistance gene, was overexpressed in Mu3_mut3 and Mu4_mut4 (McAleese 2005). Both these genes seemed likely to be involved in the observed change in tigecycline sensitivity, and the group chose these genes to investigate further. <br><br> | |||
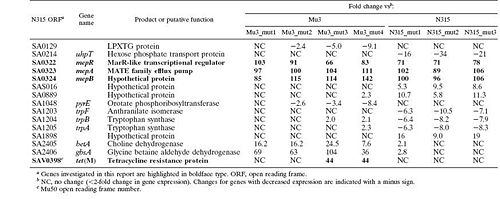
[[Image:genetable.jpg|thumb|500px|right|Figure 5. Genes with expression changes greater than 8-fold between Mu3 and N315 parent and mutant strains (McAleese 2005).]] | |||
The gene cluster ''mepRAB'' was overexpressed by 100-fold in both the Mu3 and N315 mutants. A BLAST analysis showed that ''mepR'' encodes a MarR-like transcription regulator, ''mepA'' encodes a multidrug and toxin extrusion (MATE) family efflux pump. The best characterized MATE pump is the NorM efflux pump in ''Vibrio parahaemolyticus'' which is Na<sup>+</sup>-driven. Additionally, Kyte-Doolittle hydropathy analysis of the MepA protein predicts that it has 12 transmembrane domains, which is characteristic of the MATE family pumps. The gene ''mepB'' encodes a protein of unknown function, though because of its upregulation, it seems likely that the protein will be found to play a role in tigecycline resistance. To confirm the effect of overexpression of ''mepA'', a multicopy plasmid containing the gene was inserted into the parent Mu3 and N315 strains. This resulted in a four-fold increase in the minimum inhibitory concentration (MIC) of tigecycline, a measure of resistance to the antibiotic. This suggests that the bacteria are likely using the ''mepA'' efflux pump to export tigecycline from their cells. <br><br> | |||
The ''tetM'' gene was overexpressed in two of the four Mu3 mutants. They attributed this to a 87-bp deletion in the ''tetM'' promoter region that removed a stem-loop-forming region that likely regulated the gene by allowing for transcription attenuation. The loss of this region would cause the gene to be overexpressed. Experiments similar to those used to test the effect of ''mepA'' overexpression were carried out to determine the effect of TetM on tigecycline susceptibility. They found that TetM alone was not enough to produce resistance to tigecycline. Even in combination with the MepA efflux pump, no additional resistance to tigecycline seemed to be produced by the overproduction of TetM, suggesting that even though both mutations were present in these strains, they do not seem to work synergistically (McAleese 2005). Also, this shows that tigecycline seems to be able to completely resist the effects of TetM, giving it a distinct advantage over tetracycline for use in treating infections. <br><br> | |||
<br> | |||
==Conclusion== | ==Conclusion== | ||
<br> | <br>In the face of thousands of species of pathogenic bacteria that are constantly evolving and developing new ways to resist our efforts, finding ways to fight back and treat the infections they cause can feel like an almost pointless task. It seems probable that any antibiotic we can develop will eventually lead to corresponding resistance in bacterial populations. This presents a huge problem for hospital patients and community-acquired infections. All the most widely-used antibiotics, including penicillin, methicillin, vancomycin, and tetracycline, have induced resistance in a variety of bacterial species and will likely become entirely defunct eventually. Therefore, the development of new antibiotics is a race against the evolutionary clock, a test to see if we can develop new antibiotics before bacteria become resistant to the old ones. <br><br> | ||
One promising advance for our side has been the creation of the glycylcycline class of antibiotics. The success of tigecycline, though derived from tetracycline, in evading the tetracyclin resistance mechanisms buys us at least a little more time and provides hope for more such effective antibiotics to emerge from the glycylcycline class. It's observed to bind to the ribosome in new orientation and more tightly than its predecessor, tetracycline, and as a result is a more effective antibiotic. It is also able to skirt the typical bacterial resistance mechanisms of efflux pumps and ribosomal protection proteins. Even still, laboratory-derived strains of ''S. aureus'' that are resistant to tigecycline have already been found. These strains employ novel tools to overcome the antibiotic, including the ''mepA'' MATE family efflux pump and foreshadow the time when tigecycline too will be ineffective against bacteria. By studying the way in which tigecycline achieves antimicrobial activity and trying to predict the ways in which it can and probably will be made ineffective, we can begin to better understand why it is so successful and think ahead to the next wave of antibiotics we will inevitably need to develop. | |||
<br> | |||
==References== | ==References== | ||
[ | [http://jac.oxfordjournals.org/cgi/content/full/53/4/592 Bauer, G., Berens, C., Projan, S., and Hillen, W. "Comparison of tetracycline and tigecycline binding to ribosomes mapped by dimethylsulphate and drug-directed Fe<sup>2+</sup> cleavage of 16S rRNA". ''Journal of Antimicrobial Chemotherapy''. 2004. Volume 53. p. 592-599.] | ||
[http://www.jpgmonline.com/article.asp?issn=0022-3859;year=2009;volume=55;issue=1;spage=65;epage=68;aulast=Bhattacharya Bhattacharya, M., Parakh, A., and Narang, M. "Tigecycline". ''J. Postgrad. Med.''. 2009. Volume 55. p. 65-68.] | |||
[http://www.drupjournal.com/article/S1368-7646(02)00051-1/abstract Chopra, Ian. "New developments in tetracycline antibiotics: glycylcylines and tetracycline efflux pump inhibitors". ''Drug Resistance Updates''. 2002. Volume 5. P. 119-125] | |||
[http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=1426172 Greer, N. "Tigecycline (Tygacil): the fist in the glycylcycline class of antibiotics". ''Pharmacology Notes''. 2006. Volume 19. p. 155-161.] | |||
[http://aac.asm.org/cgi/content/full/50/6/2156?maxtoshow=&HITS=10&hits=10&RESULTFORMAT=&fulltext=%B5m&searchid=1&FIRSTINDEX=2820&resourcetype=HWFIG Olson, M., Ruzin, A., Feyfant, E., Rush, T., O'Connell, J., and Bradford, P. "Functional, Biophysical, and Structural Bases for Antibacterial Activity of Tigecycline". ''Antimicrobial Agents and Chemotherapy''. 2006. Volume 50. p. 2156-2166.] | |||
[http://aac.asm.org/cgi/content/full/49/5/1865 McAleese, F., Petersen, P., Ruzin, A., Dunman, P., Murphy, E., Projan, S., and Bradford, P. "A Novel MATE Family Efflux Pump Contributes to the Reduced Susceptibility of Laboratory-Derived ''Staphylococcus aureus'' Mutants to Tigecycline". ''Antimicrobial Agents and Chemotherapy''. 2005. Volume 49. p. 1865-1871.] | |||
Edited by student of [mailto:slonczewski@kenyon.edu Joan Slonczewski] for [http://biology.kenyon.edu/courses/biol238/biol238syl09.html BIOL 238 Microbiology], 2009, [http://www.kenyon.edu/index.xml Kenyon College]. | Edited by student of [mailto:slonczewski@kenyon.edu Joan Slonczewski] for [http://biology.kenyon.edu/courses/biol238/biol238syl09.html BIOL 238 Microbiology], 2009, [http://www.kenyon.edu/index.xml Kenyon College]. | ||
Latest revision as of 20:15, 10 August 2010
By: Kristina Buschur
Introduction
The ability of bacteria to quickly develop resistance to commonly used antibiotics is a huge hurdle in the path of disease treatment. Because of this, there is an ever-present need to develop new antibiotics that use novel mechanisms to overcome multidrug-resistance and prevent microbial growth. Since tetracycline has been in wide use since the mid-1900s for treatment of many human and animal infections and as growth promoters in agriculture, many bacteria have since developed sophisticated mechanisms to prevent the harmful effects of the antibiotic (Greer 2006). Specifically, tetracycline-speicific efflux pumps and ribosomal protection proteins are commonly employed by bacteria. The glycylcycline class of antibiotics is a powerful tool that has been recently developed to combat this problem. Developed by Lederle Laboratories (now American Home Products) in the early 1990s and derived from tetracycline, glycylcyclines have added substituents that interfere with the mechanisms bacteria employ to resist tetracycline, including both the efflux pumps and ribosomal protection proteins (Chopra 2002).
Currently tigecycline (previously GAR-936) is the only antibiotic of the glycylcycline class in clinical use (Figure 1). Tigecycline was approved by the Food and Drug Administration in 2005, and is particularly useful in the treatment of multi-drug resistant infections, which are especially hard to eradicate. Penicillin-resistant Streptococcus pneumoniae, methicillin-resistant Staphylococcus aureus and Staphylococcus epidermidis, and vancomycin-resistant Enterococcus species are several examples of species that have developed an antibiotic resistance but are still affected by tigecycline (Greer 2006). In addition to these, there is a wide variety of organisms, including both gram-positive and gram-negative bacteria, that are sensitive to tigecycline. It does not, however, seem to have any effect on Proteus, Providencia, or Pseudomonas species. Even still, tigecycline shows particular success against some anaerobic organisms including Bacteriodes fragilis, Peptostreptococcus species, and Propionibacterium acnes which for the most part are able to evade tetracycline.
The tigecycline antibiotic is structurally very similar to minocycline and similarly binds to the bacterial 30S ribosome unit. The manner in which the molecule binds prevents amino-acyl tRNAs from binding to the A site of the ribosome and subsequently prevents peptide formation and bacterial growth. The main difference between tigecycline and minocycline is the addition of an N,N-dimethylglycylamido group which actually causes the molecule to bind to the ribosome up to five times more tightly which in turn decreases the probability that resistance will develop. In general, the glycylcycline class of antibiotics (the 9-glycinyltetracyclines) is characterized by a molecular structure containing a four-ring carbocyclic skeleton with a substitution of an N-alkylglycylamido group at the D-9 position (Bhattacharya 2009). Biochemical experiments have shown that the tigecycline molecule binds to the same site on 16S rRNA as tetracycline but in a different orientation and with greater affinity. This has been confirmed by experiments in which a decreased binding affinity was observed in strains of E. coli with known mutations in the A site of the rRNA (G966U or G1058C).
Tigecycline (marketed as Tygacil) is used for the treatment of complicated skin and skin structure infections and complicated intraabdominal infections caused by susceptible organisms. The drug is administered to a patient intravenously with a dose of 50 mg every 12 hours after an initial 100 mg loading dose (Greer 2006). This dosage is recommended for patients regardless of age, sex, or race. After being administered, it is found at a higher concentration in the gallbladder, lungs, and colon than in the serum. In the bone and synovial fluid, though, it is found in lower concentrations than in the serum. Its elimination half-life is about 36 hours, and it leaves the body mainly through the bile and feces, but partly through the urine as well. As with nearly all drugs, tigecycline has several known side-effects, including nausea, vomiting, and diarrhea, but these are relatively minor. Barring any severe complications, the treatment period usually ranges from five to fourteen days (Bhattacharya 2009).
Comparison of Tetracycline and Tigecycline Ribosome Binding
Bauer et al. have identified the binding sites of tetracycline and tegecycline through probing 70S E. coli ribosomes with dimethylsulphate (DMS) and Fe2+-mediated cleavage. This method works by allowing Fe2+ to replace Mg2+ complexed with tetracyclines. When H2O2 is added, the Fe2+ produces transient, very reactive hydroxy radicals that can cleave RNA close to where tetracycline is bound. It follows then, that the resulting cleavage sites will also be the location at which tetracycline binds to the ribosome (Bauer 2004). Crystallography experiments have shown that there are six tetracycline binding sites on the 30S ribosome.
Three sites in the rRNA were found that were cleaved in the presence of tetracycline. Two of the three sites, those in h31 and h34, corresponded with tetracycline binding site-1. These were confirmed through a point mutation (G1058 to C) in h34 that produced tetracycline resistance. Additional mutation experiments showed that mutations in these areas also resulted in an eight-fold increase in resistance to tigecycline. The third rRNA cleavage site in h29 corresponds to tetracycline binding site-4, so the observation is confirmed by crystallography data. There is, however, no evidence of a magnesium ion complexed with tetracycline at tetracycline binding site-2. It follows, then, that the Fe2+-mediated cleavage technique would not reveal a tetracycline at this site. The other tetracycline binding sites have previously shown no evidence of being involved in antibiotic activity.
With this knowledge, the authors conclude that since the three cleavage sites correspond to the physiologically important tetracycline binding sites and since the experiments produced identical results whether tetracycline or tigecycline was used, then tetracycline and tigecycline must share the same binding sites on 16S rRNA. Furthermore, tigecycline appears to bind much more tightly and at lower concentrations. There were, however, some differences in the Fe2+ cleavages intensities between tetracycline and tigecyclince. So although they do bind the ribosome in the same location, tigecycline likely binds in a different orientation than does tetracycline. The authors propose that the bulky t-butylglycylamido substituent at position 9 on tigecycline but absent on tetracycline is responsible for this difference. To further investigate this, they compare the superimposed co-crystal structures of tetracycline and DMG-DMDOT (a glycylcycline derivative very similar to tigecycline) complexed with helices 31 and 34, which make up tetracycline binding site-1 (Figure 2). This comparison revealed that the bulky dimethylglycylamido substituent on DMG-DMOT would clash with the phosphate backbone of G1053, forcing it to bind the ribosome differently (Bauer 2004). Likewise, the t-butylglycylamido substituent on tigecycline is even bigger and would presumably change the orientation in which tigecycline binds in a similar, if not more dramatic, way.
Antibacterial Activity of Tigecycline
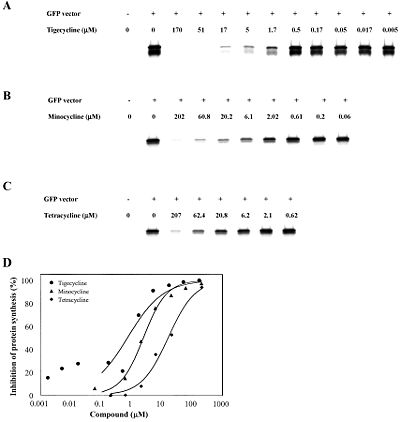
In 2006, Olson et al. reported using functional, biophysical, and molecular modeling experiments to determine the way in which tigecycline demonstrates antibacterial activity. To investigate the effect of tigecycline, in comparison with tetracycline and minocycline, on bacterial protein synthesis, they used in vitro transcription/translation (IVT) assays in which E. coli cell lysate was supplied with a GFP vector and the amount GFP produced was measured with a phosphorimager. The results clearly show a pattern in which the amount of GFP produced decreases quickly as the concentration of tigecycline, minocycline, or tetracycline is increased. This suggests that each of these molecules inhibits translation (Figure 3). Although all three antibiotics exhibit this trend, it is also important to note that tigecycline started inhibiting translation at the lowest concentration. At a tigecycline concentration of just 1.7 µM a significant inhibition of protein synthesis can be observed. This is in comparison to the much higher concentrations of minocycline (6.1 µM) and tetracycline (62.4 µM) that are required before any notable inhibition can be observed. These observations are summarized in Figure 3d, which presents a half maximal inhibitory concentration (IC50) anaylsis. The graph makes it clear that over a large range of concentrations, tigecycline has a much higher percent inhibition of protein synthesis than minocycline, which in turn is also more effective than tetracycline. This pattern also makes sense when it is considered that minocycline is a tetracyline derivative, and tigecycline is a minocycline derivative (Figure 4).
To further investigate the potency of these three drugs, competition studies very performed. They observed which of the antibiotics bound most effectively to both the 30S and the 70S subunits, and it was found that tigecycline bound to both the 30S and 70S subunits with 5-fold greater affinity than minocycline and 100-fold greater affinity than tetracycline (Olson 2006). Again, this trend is consistent with the structures of these molecules.
Finally, computational docking studies were performed to investigate the tigecycline-ribosome interaction. One particular position showed energetically favorable interactions that were consistent with all previous data. This position demonstrated favorable electrostatic, van der Waals, and hydrogen-bonding interactions with the 30S subunit that suggested it associated with the ribosome more closely than both minocycline and tetracycline. The overall orientation was very similar to that previously described for tetracycline, with the Mg ion interacting with the phosphate groups of G1197, C1054, and U1196. The most notable difference, though, was that the glycylcyclines interacted directly with other areas of the A site. These interactions included additonal hydrogen-bonding and van der Waals interactions that had never been seen before in that area (Olson 2006), observations that are consistent with the increased binding affinity of the glycylcyline class of antibiotics.
Combined, these observations further support the conclusion of Bauer et al. that tigecycline sterically inhibits the binding of aminoacyl-tRNA to the A site of the ribosome in a manner very similar to that of tetracycline. Furthermore, their data suggest that the way in which glycylcyclines are able to avoid TetM-mediated resistance could be the result of either their increased affinity for binding the ribosome or the unique orientation in which they bind the ribosome that might make TetM ineffective in protecting the ribosome. Perhaps, even, it is some combination of these two qualities that makes glycylcylines so effective.
Tigecycline Resistance Mechanisms
Despite its current success in preventing growth of bacterial species that have already developed resistance to tetracycline, it is inevitable that bacteria will eventually mutate so that they are resistant to tigecycline as well. This is particularly a concern since tigecycline is so structurally similar to tetracycline, an antibiotic many bacteria have already developed successful defenses against. It is thought that resistance to tigecycline will develop even more quickly as a result of this (Chopra 2002). McAleese et al. have already found laboratory-derived mutants of Staphylococcus aureus that exhibit resistance to tigecycline and have traced that resistance to increased expression of the multidrug export protein mepRAB gene cluster. This cluster includes mepA, a novel MATE family efflux pump.
While there are no known naturally occurring isolates of S. aureus that show resistance to tigecycline, two such mutants were found by serial passages through increasing concentrations of the antibiotic. One mutant was derived from the Mu3 strain (a clinical MRSA isolate with heterogeneous resistance to vancomycin) and the other from the N315 strain (a clinical MRSA isolate). The bacteria that resulted from this procedure had a very much reduced sensitivity to tigecycline. A susceptibility profile of the mutants was performed to determine if the sensitivity to other antibiotics had decreased as well, which might suggest a more general resistance mechanism was at work. They found that both of the mutants were more resistant to DMG-MINO and DMG-DMDOT, two other glycylcylcline antibiotics very similar in structure to tigecycline. Interestingly, the N315 mutant also showed increased resistance to EtBr and TPP (McAleese 2005).
A transcriptional profile analysis was undertaken to determine the origin of the decreased susceptibility observed in these mutants. S. aureus GeneChips were used to compare the DNA and RNA profiles of the Mu3 and N315 with their tigecycline-resistant mutants. They looked for changes in DNA content or gene expression that could explain the increased resistance. The found no changes in the DNA present in the mutants from the parent strains, suggesting that the mutants hadn't lost any genes that might be causing them to be susceptible to the antibiotic. The technique, though, was not sensitive enough to detect point mutations or small deletions, so it is still possible that these changes had occured. RNA was harvested from the parent and mutant strains during mid-exponential growth to compare their transcriptional profiles. Fifteen genes were found (though some of them are only hypothetical) whose transcription saw at least an eight-fold difference between the parent and the mutant (Figure 5). Several of these genes that are believed to code hypothetical proteins and other genes provided functions for which a connection to tigecycline resistance was not immediately apparent. Among these fifteen genes, though, was mepA, an efflux pump-encoding gene. Additionally, tetM, a tetracycline resistance gene, was overexpressed in Mu3_mut3 and Mu4_mut4 (McAleese 2005). Both these genes seemed likely to be involved in the observed change in tigecycline sensitivity, and the group chose these genes to investigate further.
The gene cluster mepRAB was overexpressed by 100-fold in both the Mu3 and N315 mutants. A BLAST analysis showed that mepR encodes a MarR-like transcription regulator, mepA encodes a multidrug and toxin extrusion (MATE) family efflux pump. The best characterized MATE pump is the NorM efflux pump in Vibrio parahaemolyticus which is Na+-driven. Additionally, Kyte-Doolittle hydropathy analysis of the MepA protein predicts that it has 12 transmembrane domains, which is characteristic of the MATE family pumps. The gene mepB encodes a protein of unknown function, though because of its upregulation, it seems likely that the protein will be found to play a role in tigecycline resistance. To confirm the effect of overexpression of mepA, a multicopy plasmid containing the gene was inserted into the parent Mu3 and N315 strains. This resulted in a four-fold increase in the minimum inhibitory concentration (MIC) of tigecycline, a measure of resistance to the antibiotic. This suggests that the bacteria are likely using the mepA efflux pump to export tigecycline from their cells.
The tetM gene was overexpressed in two of the four Mu3 mutants. They attributed this to a 87-bp deletion in the tetM promoter region that removed a stem-loop-forming region that likely regulated the gene by allowing for transcription attenuation. The loss of this region would cause the gene to be overexpressed. Experiments similar to those used to test the effect of mepA overexpression were carried out to determine the effect of TetM on tigecycline susceptibility. They found that TetM alone was not enough to produce resistance to tigecycline. Even in combination with the MepA efflux pump, no additional resistance to tigecycline seemed to be produced by the overproduction of TetM, suggesting that even though both mutations were present in these strains, they do not seem to work synergistically (McAleese 2005). Also, this shows that tigecycline seems to be able to completely resist the effects of TetM, giving it a distinct advantage over tetracycline for use in treating infections.
Conclusion
In the face of thousands of species of pathogenic bacteria that are constantly evolving and developing new ways to resist our efforts, finding ways to fight back and treat the infections they cause can feel like an almost pointless task. It seems probable that any antibiotic we can develop will eventually lead to corresponding resistance in bacterial populations. This presents a huge problem for hospital patients and community-acquired infections. All the most widely-used antibiotics, including penicillin, methicillin, vancomycin, and tetracycline, have induced resistance in a variety of bacterial species and will likely become entirely defunct eventually. Therefore, the development of new antibiotics is a race against the evolutionary clock, a test to see if we can develop new antibiotics before bacteria become resistant to the old ones.
One promising advance for our side has been the creation of the glycylcycline class of antibiotics. The success of tigecycline, though derived from tetracycline, in evading the tetracyclin resistance mechanisms buys us at least a little more time and provides hope for more such effective antibiotics to emerge from the glycylcycline class. It's observed to bind to the ribosome in new orientation and more tightly than its predecessor, tetracycline, and as a result is a more effective antibiotic. It is also able to skirt the typical bacterial resistance mechanisms of efflux pumps and ribosomal protection proteins. Even still, laboratory-derived strains of S. aureus that are resistant to tigecycline have already been found. These strains employ novel tools to overcome the antibiotic, including the mepA MATE family efflux pump and foreshadow the time when tigecycline too will be ineffective against bacteria. By studying the way in which tigecycline achieves antimicrobial activity and trying to predict the ways in which it can and probably will be made ineffective, we can begin to better understand why it is so successful and think ahead to the next wave of antibiotics we will inevitably need to develop.
References
Edited by student of Joan Slonczewski for BIOL 238 Microbiology, 2009, Kenyon College.