Rhodococcus ruber: Difference between revisions
| Line 24: | Line 24: | ||
==Description and Significance== | ==Description and Significance== | ||
Historically, the genus ''Rhodococcus'' was first defined by Zopf in 1891. ''Nocardia rubra'', or what is known today as ''Rhodococcus ruber'', was isolated from a soil sludge by Kruse in 1896. However, there were issues with classification due to morphological and staining identification techniques. In 1977, Goodfellow and Alderson did a complete reclassification of the genus ''Rhodococcus'', which resulted in ''Norcordia rubra'' being amended as ''Rhodococcus ruber''. | |||
The term ''Rhodococcus'' is from the combination of the Greek words “rhodon” and “coccus” meaning “the rose” and “the grain“ respectively. The morphological structure of ''R. ruber'' first starts as long rods, then breaks off into short rods and cocci throughout different growth phases. ''Rhodococcus ruber'' is a gram positive, non-motile, non-spore forming bacteria. | |||
==Genome Structure== | ==Genome Structure== | ||
Revision as of 07:02, 29 April 2020
Classification
Domain: Bacteria
Phylum: Actinobacteria
Class: Actinobacteria
Order: Corynbacteriales
Family: Nocardiaceae
Genus: Rhodococcus
Species
|
NCBI: [1] |
Rhodococcus ruber
Description and Significance
Historically, the genus Rhodococcus was first defined by Zopf in 1891. Nocardia rubra, or what is known today as Rhodococcus ruber, was isolated from a soil sludge by Kruse in 1896. However, there were issues with classification due to morphological and staining identification techniques. In 1977, Goodfellow and Alderson did a complete reclassification of the genus Rhodococcus, which resulted in Norcordia rubra being amended as Rhodococcus ruber.
The term Rhodococcus is from the combination of the Greek words “rhodon” and “coccus” meaning “the rose” and “the grain“ respectively. The morphological structure of R. ruber first starts as long rods, then breaks off into short rods and cocci throughout different growth phases. Rhodococcus ruber is a gram positive, non-motile, non-spore forming bacteria.
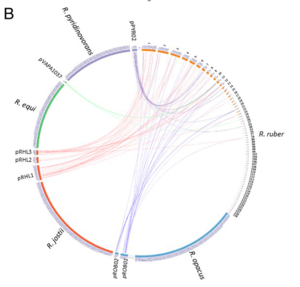
Genome Structure
12 genome studies have been reported for the following strains of Rhodococcus ruber: R. ruber str. YC-YT1, R. ruber str. YYL, R. ruber str. P14, R. ruber str. SD3, R. ruber str. R1, R. ruber str. DSM 43338, R. ruber str. P25, R. tuber IEGM 231, R. ruber str. OA1, R. ruber str. BKS 20-38, R. ruber str. Chol-4, and R. ruber str. NCRX 15591.
The genome size of Rhodococcus ruber is on average 5.57 Mb. The genome size varies in some strains and larger genome sizes can be attributed to the presence of plasmids. The strains of R. ruber that contain plasmids are YC-TC1, YYL, and R1. The average amount of protein coding genes is 4932 and the average G+C content is 70.49%. A high guanine-cytosine content is characteristic of Rhodococcus species.
Cell Structure, Metabolism and Life Cycle
Interesting features of cell structure; how it gains energy; what important molecules it produces.
Ecology and Pathogenesis
Habitat; symbiosis; biogeochemical significance; contributions to environment.
If relevant, how does this organism cause disease? Human, animal, plant hosts? Virulence factors, as well as patient symptoms.
References
[2] Alvarez, H. M. Biology of Rhodococcus; Springer, 2010; Vol. 16.
[3] Chae, Y.; An, Y.-J. Current research trends on plastic pollution and ecological impacts on the soil ecosystem: A review. Environmental Pollution 2018, 240, 387–395.
[4] Finnerty, W. R. The Biology and Genetics of the Genus Rhodococcus. Annual Review of Microbiology 1992, 46, 193–218.
[5] Gravouil, K.; Ferru-Clément, R.; Colas, S.; Helye, R.; Kadri, L.; Bourdeau, L.; Moumen, B.; Mercier, A.; Ferreira, T. Transcriptomics and Lipidomics of the Environmental Strain Rhodococcus ruber Point out Consumption Pathways and Potential Metabolic Bottlenecks for Polyethylene Degradation. Environmental Science & Technology 2017, 51, 5172–5181.
[6] Gilan (Orr), I.; Hadar, Y.; Sivan, A. Colonization, biofilm formation and biodegradation of polyethylene by a strain of Rhodococcus ruber. Applied Microbiology and Biotechnology 2004, 65.
[7] Heras, L. F. D. L.; Fernández, E. G.; Llorens, J. M. N.; Perera, J.; Drzyzga, O. Morphological, Physiological, and Molecular Characterization of a Newly Isolated Steroid-Degrading Actinomycete, Identified as Rhodococcus ruber Strain Chol-4. Current Microbiology 2009, 59, 548–553.
[8] Montazer, Z.; Najafi, M. B. H.; Levin, D. B. Challenges with Verifying Microbial Degradation of Polyethylene. Polymers 2020, 12.
[9] Mor, R.; Sivan, A. Biofilm formation and partial biodegradation of polystyrene by the actinomycete Rhodococcus ruber. Biodegradation 2008, 19, 851–858.
[10] Santo, M.; Weitsman, R.; Sivan, A. The role of the copper-binding enzyme – laccase – in the biodegradation of polyethylene by the actinomycete Rhodococcus ruber. International Biodeterioration & Biodegradation 2013, 84, 204–210.
[11] Sivan, A.; Szanto, M.; Pavlov, V. Biofilm development of the polyethylene-degrading bacterium Rhodococcus ruber. Applied Microbiology and Biotechnology 2006, 72, 346–352.
Author
Page authored by Hannah von Werder, student of Prof. Jay Lennon at Indiana University.