CTXφ Bacteriophage: Difference between revisions
| Line 11: | Line 11: | ||
==Infection, Replication & Release from Host Cell== | ==Infection, Replication & Release from Host Cell== | ||
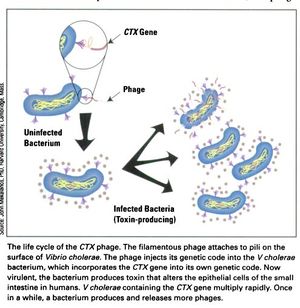
[[Image:CTX_Phage_life_cycle.jpg|thumb|right|<b>Figure 3:</b> The life cycle of the CTXφ Bacteriophage with <i>Vibrio cholerae</i> as its host.]] | [[Image:CTX_Phage_life_cycle.jpg|thumb|right|<b>Figure 3:</b> The life cycle of the CTXφ Bacteriophage with <i>Vibrio cholerae</i> as its host.]] | ||
In a process known as lysogenic phage conversion, the CTXφ bacteriophage integrates, among others, its <i>ctxAB</i> genes into its host, <i>Vibrio cholerae</i>. The <i>ctxAB</i> genes code for a type of exotoxin, called Cholera toxin or just "CT," that causes <i> V. cholerae</i> to switch from being nonpathogenic to highly virulent. For those infected, it is also the primary cause of the large amounts of diarrhea—cholera's main symptom.<ref name = Boyd>Boyd, E. F. (2010). "Efficiency and specificity of CTXphi chromosomal integration: dif makes all the difference." Proceedings of the National Academy of Sciences of the United States of America, 107(9), 3951–3952. https://doi.org/10.1073/pnas.1000310107</ref> After finding a live host and undergoing CTXφ particle adsorption (see Figure 3 at right) to the <i>V. cholerae</i> cell wall, viral ssDNA is then injected into the cell cytoplasm and immediately forms a circular plasmid, pCTX, which then integrates into the <i>V. cholerae</i> genome at a site-specific attachment location.<ref name = Boyd>Boyd, E. F. (2010). "Efficiency and specificity of CTXphi chromosomal integration: dif makes all the difference." Proceedings of the National Academy of Sciences of the United States of America, 107(9), 3951–3952. https://doi.org/10.1073/pnas.1000310107</ref> Because the CTXφ phage, like all bacteriophages, only injects its genome, or, more commonly, <i>prophage</i>, the CTXφ integration makes use of many of the host's enzymes. | In a process known as lysogenic phage conversion, the CTXφ bacteriophage integrates, among others, its <i>ctxAB</i> genes into its host, <i>Vibrio cholerae</i>. The <i>ctxAB</i> genes code for a type of exotoxin, called Cholera toxin or just "CT," that causes <i> V. cholerae</i> to switch from being nonpathogenic to highly virulent. For those infected, it is also the primary cause of the large amounts of diarrhea—cholera's main symptom.<ref name = Boyd>Boyd, E. F. (2010). "Efficiency and specificity of CTXphi chromosomal integration: dif makes all the difference." Proceedings of the National Academy of Sciences of the United States of America, 107(9), 3951–3952. https://doi.org/10.1073/pnas.1000310107</ref> After finding a live host and undergoing CTXφ particle adsorption (see Figure 3 at right) to the <i>V. cholerae</i> cell wall, viral ssDNA is then injected into the cell cytoplasm and immediately forms a circular plasmid, pCTX, which then integrates into the <i>V. cholerae</i> genome at a site-specific attachment location.<ref name = Boyd>Boyd, E. F. (2010). "Efficiency and specificity of CTXphi chromosomal integration: dif makes all the difference." Proceedings of the National Academy of Sciences of the United States of America, 107(9), 3951–3952. https://doi.org/10.1073/pnas.1000310107</ref> Because the CTXφ phage, like all bacteriophages, only injects its genome, or, more commonly, its <i>prophage</i>, the CTXφ integration makes use of many of the host's enzymes. | ||
Revision as of 02:05, 12 December 2020
Overview
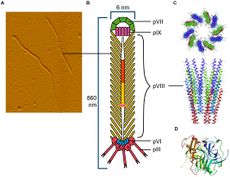
The CTXφ bacteriophage (or sometimes written as CTXphi bacteriophage) is a lysogenic, filamentous, single-stranded DNA (ssDNA) phage that is responsible for turning the previously non-infectious Vibrio cholerae into a highly pathogenic microbe that causes disease in humans.[2][3][4] Vibrio virus CTXphi belongs to the family Inoviridae and the genus Affertcholeramvirus. These phages are comprised of only 5 viral proteins and are between 1 and 2 microns in length.[5] They resemble something like a cooked spaghetti noodle. (See Figure 1 at right).
Genetic Material
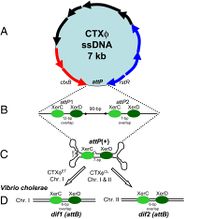
The CTXφ phage can be identified in its host in both its replicative form (when the bacteriophage is in lysogeny, or the lytic cycle), and in its non-replicative form, which is most common. This phage, when in its lysogenic cycle, integrates its own genetic material in such a way as to form the most stable lysogens.[4]
Infection, Replication & Release from Host Cell
In a process known as lysogenic phage conversion, the CTXφ bacteriophage integrates, among others, its ctxAB genes into its host, Vibrio cholerae. The ctxAB genes code for a type of exotoxin, called Cholera toxin or just "CT," that causes V. cholerae to switch from being nonpathogenic to highly virulent. For those infected, it is also the primary cause of the large amounts of diarrhea—cholera's main symptom.[4] After finding a live host and undergoing CTXφ particle adsorption (see Figure 3 at right) to the V. cholerae cell wall, viral ssDNA is then injected into the cell cytoplasm and immediately forms a circular plasmid, pCTX, which then integrates into the V. cholerae genome at a site-specific attachment location.[4] Because the CTXφ phage, like all bacteriophages, only injects its genome, or, more commonly, its prophage, the CTXφ integration makes use of many of the host's enzymes.
Other examples:
Bold
Italic
Subscript: H2O
Superscript: Fe3+
CT & non-CT Toxins
The CTXφ phage contains both types of exotoxins.
CT Toxins
Non-CT Toxins
Conclusion
Overall text length should be at least 1,000 words (before counting references), with at least 2 images. Include at least 5 references under Reference section.
References
- ↑ Gagic, D., Ciric M., Wen W., Ng F., Rakonjac J. (2016). "Exploring the Secretomes of Microbes and Microbial Communities Using Filamentous Phage Display." Frontiers in Microbiology, 7:429. https://doi.org/10.3389/fmicb.2016.00429.
- ↑ Davis, B. M., Kimsey, H. H., Chang, W., & Waldor, M. K. (1999). "The Vibrio cholerae O139 Calcutta bacteriophage CTXφ is infectious and encodes a novel repressor." Journal of Bacteriology, 181(21), 6779-6787. https://www.frontiersin.org/articles/10.3389/fmicb.2016.00429/full
- ↑ Ochman, H., Lawrence, J. & Groisman, E. (2000). "Lateral gene transfer and the nature of bacterial innovation." Nature, 405, 299–304. https://doi.org/10.1038/35012500.
- ↑ 4.0 4.1 4.2 4.3 4.4 Boyd, E. F. (2010). "Efficiency and specificity of CTXphi chromosomal integration: dif makes all the difference." Proceedings of the National Academy of Sciences of the United States of America, 107(9), 3951–3952. https://doi.org/10.1073/pnas.1000310107
- ↑ Mai-Prochnow, A., Hui, J., Kjelleberg, S., Rakonjac, J., McDougald, D., Rice, S. (2015). "Big things in small packages: the genetics of filamentous phage and effects on fitness of their host." FEMS Microbiology Reviews, Volume 39, Issue 4, Pages 465–487, https://doi.org/10.1093/femsre/fuu007
Edited by Tara Cerny, student of Joan Slonczewski for BIOL 116 Information in Living Systems, 2019, Kenyon College.