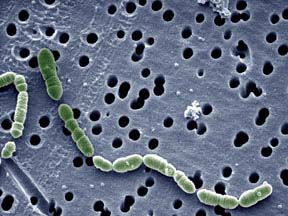
Oenococcus oeni
ated}}
Classification
Domain: Bacteria
Phylum: Firmicutes
Class: Bacilli
Order: Lactobacillales
Family: Leuconostocaceae
Genus: Oenococcus
Species: oeni (1)
NCBI link to find]
Species
|
NCBI: Taxonomy |
Oenococcus oeni
Description and Significance
Oenoccoccus oeni is a gram-positive, nonmotile, chemoorganotrophic lactic acid bacteria that grows in chains of circular to ellipsoidal cells (2). Unlike many bacteria, it thrives in the harsh environment of wine and is considered a beneficial to wine quality and character, rather an agent of damaging value (3). O. oeni is beneficial to wine and the entire field of oenology because of its primary ability to perform malolactic fermentation Describe the appearance, habitat, etc. of the organism, and why you think it is important.
Genome Structure
Any strain of O. onei has about 1800 protein-coding genes Describe the size and content of the genome. How many chromosomes? Circular or linear? Other interesting features? What is known about its sequence?
Cell Structure, Metabolism and Life Cycle
Interesting features of cell structure; how it gains energy; what important molecules it produces.
Ecology and Pathogenesis
Habitat; symbiosis; biogeochemical significance; contributions to environment.
If relevant, how does this organism cause disease? Human, animal, plant hosts? Virulence factors, as well as patient symptoms.
References
(1) NCBI Genome Project [1] (2) [Dicks, L. M. T., Dellaglio, F., Collins, M. D. “Proposal To Reclassify Leuconostoc oenos as Oenococcus oeni [corrig.] gen. nov., comb. nov”. “International Journal of Systematic Bacteriology”. 1995. Volume 445. P. 395-397] (3) Borneman, A. R., McCarthy, J. M., Chambers, P. J., Bartowsky, E. J. “Comparative analysis of the Oenococcus oeni pan genome reveals genetic diversity in industrially-relevant pathways”. (4) Solieri, L., Genova, F., De Palola, M., Giudici, P. “Characterization and technological properties of Oenococcus oeni strains from wine spontaneous malolactic fermentations: a framework for selection of new starter cultures”. “Journal of Applied Microbiology”. 2010. Volume 108. P. 285-298
Author
Page authored by Drew Wehrle student of Prof. Jay Lennon at Indiana University.