Ciliophora
A Microbial Biorealm page on the Ciliophora
|
NCBI: |
Classification
Higher order taxa:
Eukaryota; Alveolata
Species:
Spirostomum minus, Zoothamnium pararbuscula, Paramecium tetraurelia
Description and Significance
There are roughly 8,000 species of Ciliophora. Ji et. al. (2005) recently discovered two new speices of Ciliophora: Pseudovorticella clampi and Zoothamnium pararbuscula. There are numerous types of Ciliophora. Some of the major ones include Didinium, Paramecium, Stentor,Suctoria, and Vorticella
Genome Structure
Some Ciliophora have unique genetic codes. In certain species, the UGA, UAG, and UAA codons, which are typically stop codons, have different functions. UAA and UAG instead code for glutamine, while UGA (the universal stop codon) translates to cysteine or tryptophan. The research of Kim et. al. (2005) suggests that this codon reassignment is mainly influenced by Eukaryotic release factor 1 (eRF1), which is an important protien for stop codon recognition. Their work also shows that the ciliate genetic code has deviated from the universal genetic code.
Cell Structure and Metabolism
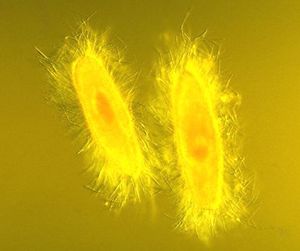
Ciliophora get their name based on their method of locomotion: they swim with cilia. Cilia are short, hairlike projections of cytoplasm composed of pairs of microtubules surrounded by cell membrane. They line the cell membrane. Cilia can also be used for obtaining food. In some species, they are fused into sheets, making them efficient at sweeping up food.
The macronuclei of Ciliophora contain numerous endocytobionts. Ciliophora are multinucleate organisms. The macronucleus controls cell functions and asexual reproduction. The micronucleus is also involved with sexual reproduction.
Ciliophora are heterotrophic organisms. Some species prey on bacteria, while others eat algae, other Ciliophora, or detritus.
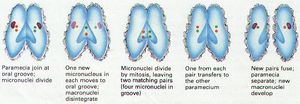
Ciliophora can reproduce sexually or asexually. Asexual reproduction by fission is the most common. Genetic exchange during sexual reproduction involves the micronucleus. After conjugation, Ciliophora divide; this produces four identical organisms.
Ecology
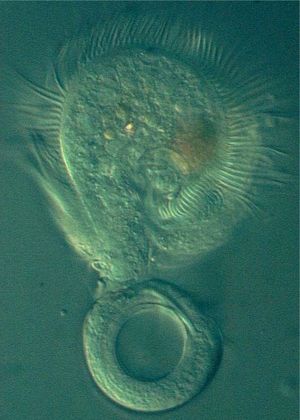
Ciliophora are mainly freshwater organisms. Ciliophora often form relationships with bacteria. Some of these may be harmful to the Ciliophora, but others are not destructive. These relationships can help increase the environmental resiliency of the bacteria. In addition Ciliophora may benefit from these relationships. The research of Fujishima et. al. (2005) illustrates that the relationship Paramecium caudatum forms with the bacterium Holospora obtusa helps the ciliate develop heat-shock resistance. Some species of Ciliophora are parasitic towards humans and other animals. Some Ciliophora species, such as Stentors, form symbiotic relationships with algae, giving them a green tint.
Some species of Ciliophora are pathogenic. Anophryoides haemophila, for example, causes Bumber Car Disease, a serious affliction of captive lobsters. It is a major cause of death among lobsters being held for commerical purposes. Bumper Car Disease results in the depletion of blood cells. There is currently no treatment for this disease.
Ciliophora are model organisms. For example, it is believed their relationships with bacteria mirror human relationships with bacteria. Brandl et. al. (2005) have studied the affects of bacteria on Ciliophora to determine its potential affects on human health.
References
Johnson, Jerry G. The World of Biology.
Lynn, D.H. 2003. The Ciliate Resource Archive. Accessed 2005 June 17.
Patterson, David J. "Parasitic Protists." The Tree of Life Web Project.
