Methanogens
A Microbial Biorealm page on the Methanogens
Methanococcus
Methanoculleus
Methanofollis
Methanopyrus
Methanosarcina
Methanosphaera
Methanothermobacter
Introduction
In 1776, Alessandro Volta ran some experiments on combustible air that was reported to him by a friend, Father Carlo Campi. On a little boat in Lake Maggiore he started to poke and stir the bottom of an area covered with reeds. Upon doing this, Volta noticed a lot of air emerging and decided to collect some in a large glass container. Upon analysis of the air he noted that it burned a beautiful blue flame. It wasn't for nearly a century that firm evidence was collected that showed that the methane formation in these habitats was a microbial process.
The discovery of methanogens helped produce the idea for the kingdom Archaeobacteria, that would include methanogens, some extreme halophiles, and some extreme thermophilic sulfur-dependant organisms. Woese et al. (1990) proposed that a urkingdom be made for the methanogens and other Archaeobacteria and it be called Archaea. Present day there is a superkingdom Archaea that contains phylums, with the two most prominent being Euryarchaeota and Crenarchaeota and the methanogens being under Euryarchaeota.
Methanogens can be used to produce methane (aka natural gas, biogas) from biomass and degrade and detoxify agricultural, municipal, and industrial wastes. Methane is also a big contributor to global warming, which is why a better understanding of how methanogenesis works is needed so that we can use methane as a renewable source of energy and limit its greenhouse gas effect.
Metabolism
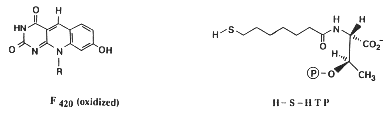
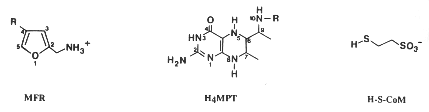
Most methanogens can grow on CO2 and H2 as their sole energy source. There are a few exceptions that only metabolize [apathway.GIF acetate], or reduce [mpathway.gif methanol] with H2, or use methylamine and methanol. For the majority that reduces [cpathway.gif CO2 to CH4], there are a few key coenzymes they need; coenzyme bound C1-intermediates Methanofuran (MFR), tetrahydromethanopterin (H4MPT), and coenzyme M (H-S-CoM). Other key coenzymes worth noting are F420 and N-7-mercaptoheptanoyl-O-phospho-L-threonine (H-S-HTP). Coenzyme F420 acts analogously to a quinone in electron transfer sequences by accepting the H+ ions from the electron donor and supplying them to the electron acceptor. The other coenzyme, H-S-HTP, does the same task as F420 only in the last step of methanogenesis from CO2 and H2.
These coenzymes are used in the most conventional form of methanogenesis, but that's not necessarily their sole purpose nor the only enzymes able to perform this task. Methanofuran is found in all methanogens but is also found in Archaeoglobus fulgidus. Tetrahydromethanopterin is also found in other archaeons besides methanogens. Not only that, but Methanosarcina spp. use tetrahydrosarcinaopterin intead, which differs from tetrahydromethanopterin by the addition of a glutamyl moiety in the substituent R). Coenzyme M, on the other hand, is found only in methanogens.
Below is a table including energy-yielding reactions used by methylotrophic methanogens. One of the more common reactions, and the reaction showed above, is the 9th one down. While the last reaction involving methane is the most favorable, methane is not as readily available as carbon dioxide.
| Reaction | δ Go (kJ/mol CH4) | |
| 1. 4 CH3OH |
→ 3 CH4 + CO2 + 2 H20 |
-106 |
| 2. CH3OH +H2 |
→ CH4 + H2O |
-112.5 |
| 3. 4 CH3NH2 + 2 H2O |
→ 3 CH4 + CO2 + 4 NH3 |
-76.7 |
| 4. 2 (CH3)2NH + 2 H2O |
→ 3 CH4 + CO2 + 2 NH3 |
-74.8 |
| 5. 4 (CH3)3N + 6 H2O |
→ 9 CH4 + 3 CO2 + 4 NH3 |
-75.8 |
| 6. 2 (CH3)2S + 2 H2O |
→ 3 CH4 + CO2 + 2 H2S |
-52.1 |
| 7. 4 (CH3)SH + 2 H20 |
→ 3 CH4 + CO2 + 4 H2S |
-51 |
| 8. (CH3)SH + H2 |
→ CH4 +H2S |
-69.3 |
| 9. 4 H2 + CO2 |
→ CH4 + 2 H2O |
-130.4 |
| 10. CH3COO- + H+ |
→ CH4 + CO2 |
-36 |
| 11. 4 CO + 2 H20 |
→ CH4 + 3 CO2 |
-211 |
Cell Structure
Archaea are cells that have a wide variety of shapes, sizes, and ultrastructural variations, not unlike Bacteria cells. Two shapes, rods and coccoid, seem to dominate the methanogens. Some examples of rod shaped cells include Methanobacterium spp. and Methanopyrus kandleri. Examples of the coccoid methanogens include species from Methanococcus and Methanosphaera to name a few. Methanoculleus and Methanogenium are coccoid as well but are irregularly shaped, possibly due to S-layers not being so strongly bonded like other wall structures. Methanogens are not just limited to these shapes, but include a plate shaped genus Methanoplanus, Methanospirillum that are long thin spirals, and Methanosarcina that are cluster of round cells.
It is recognized that methanogens lack murein typical of bacteria (eubacteria), but some contain pseudomurein, which can only be distinguished from its bacteria counterpart through chemical analysis. Those methanogens that do not possess pseudomurein have at least one paracrystalline array (S-layer). An S-layer is made up of proteins that fit together in an array like jigsaw pieces that do not covalently bind to one another, in contrast to a cell wall that is one giant covalent bond. Also, some methanogens have S-layer proteins that are glycosylated, which could increase stability, while others don't (e.g., Methanococcus spp).
Ecology
Methanogens are a diverse group of organisms that can live in a wide range of environments. They have been found in a range of salinity from freshwater to hypersaline. There are many freshwater and marine methanogens, but not many hyperhalophilic ones. The genus Methanohalophilus contain the most halophilic methanogens, who share some similarities to the aerobic archaeobacteria Halobacterium. These methylotrophic methanogens, which all belong to the Methanosarcinaceae, may have shared a common ancestor with the halobacteria since 16S and 23S rRNA sequence comparisons show a specific relationship between the two. In addition, the Methanosarcinacae are the only methanogens to possess cytochromes (possibly due to lateral gene transfer), which are found in halobacteria.
Methanogenesis is greatly limited when the surrounding temperature is below 15oC, but can still occur readily at temperatures near 100oC. Methanogens are found in both of these environments, however. Methanoculleus submarinus is found at temperatures near 15oC, while Methanopyrus kandleri is found in temperatures at the other end of the spectrum. Many other methanogens are thermophilic including Methanococcus jannaschii, growing around 85oC, and Methanothermobacter thermoautotrophicum.
With organisms growing at such high temperatures it would be logical to guess that they might have a high G + C content, but that would be incorrect. What has been hypothesized is that some of these methanogens like M. kandleri may have a protein similar to HMf, which has been found to bind to DNA nonspecifically and increase the melting point. Also, a new DNA topoisomerase, called reverse gyrase, has been found in hyperthermophilic archaeons as well as in hyperthermophilic bacteria. This enzyme produces positive supercoils in DNA that could render more tightly coiled DNA and greater stability.
Although most methanogens have a pH optima near neutral, there are some methanogens that live in extreme pH environments. Methanogenesis has been shown to occur at low pH's (pH=3.0), but the optimal pH value is near 6.0. According to Williams and Crawford (1985, cited in Ferry), a hydrogenotrophic methanogen, most likely Methanobacterium, was isolated from peat bogs and was able to grow at pH's as low as 5 while still producing some methane down to pH 3.0. There are also some alkaliphilic methanogens but they are not very extreme. One in particular, Methanohalophilus zhilinae, has been found in a hypersaline lake in Egypt which also has an optimum pH value of 9.2.
References
Ferry, James editor. Methanogenesis: Ecology, Physiology, Biochemistry, & Genetics. Chapman & Hall Inc, New York, 1993.