Nostoc azollae: Difference between revisions
| Line 23: | Line 23: | ||
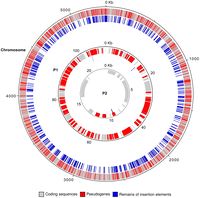
[[File:noazgenome.jpg|200px|thumb|right|Figure 1: Map of the main chromosome and plasmids of N. Azollae (9)]] | [[File:noazgenome.jpg|200px|thumb|right|Figure 1: Map of the main chromosome and plasmids of N. Azollae (9)]] | ||
==Genome Structure== | ==Genome Structure== | ||
Only a few strains of ''N. Azollae'' have been sequences, chiefly ''Nostoc Azollae 708'' (Figure 1), which had a genome size of 5.84 megabytes (8). ''N. Azollae'' possessed a small G+C content of only 38.3%, and carried with it 4 rRNA clusters alongside 44 tRNA species, making up the full set of amino acids. Notable within the gene is that intact genes, as opposed to pseudogenes, compose 52% of the genome, which is the lowest of any cyanobacteria sequenced thus far, owing partly to the massive amount of pseudogenes that comprise the rest of the genome. In addition, ''N. Azollae'' has the some of the lowest intact CDS counts of any similar cyanobacteria sequenced. | Only a few strains of ''N. Azollae'' have been sequences, chiefly ''Nostoc Azollae 708'' (Figure 1), which had a genome size of 5.84 megabytes (8). ''N. Azollae'' possessed a small G+C content of only 38.3%, and carried with it 4 rRNA clusters alongside 44 tRNA species, making up the full set of amino acids. Notable within the gene is that intact genes, as opposed to pseudogenes, compose 52% of the genome, which is the lowest of any cyanobacteria sequenced thus far, owing partly to the massive amount of pseudogenes that comprise the rest of the genome. In addition, ''N. Azollae'' has the some of the lowest intact CDS counts of any similar cyanobacteria sequenced (9). | ||
==Cell Structure, Metabolism and Life Cycle== | ==Cell Structure, Metabolism and Life Cycle== | ||
Revision as of 20:04, 29 April 2020
Classification
Domain: Bacteria
Phylum: Cyanobacteria
Class: Cyanophyceae
Order: Nostocales
Family: Nostocaceae
Genus: Nostoc
Species
|
NCBI: Taxonomy |
Nostoc azollae
Description and Significance
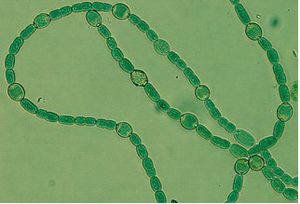
Nostoc azollae, from the genus "Nostoc", is a cyanobacteria (1), meaning it is a phototrophic organism that creates its own energy through photosynthesis. The microbe grows as unbranched filaments and differentiates itself into three seperate types of cells. Heterocysts, which differentiate in response to nitrogen deficiencies, and act as sites of fixation for the environment. These make up anywhere from 3-10% of the microbe's cells. N. azollae is also capable of creating two other kinds of cells, hormongia, consisting of short filaments, and spore-like akinetes to make up for any limitations in the environment (2).
The significance of the microbe is found within its symbiosis with Azolla filiculoides, a type of plant native to warm tropical regions all across the globe (3). The fern possesses numerous small cavities all along its surface that house N. azollae and allow for the transfer of nutrients between the two symbionts (4).
Genome Structure
Only a few strains of N. Azollae have been sequences, chiefly Nostoc Azollae 708 (Figure 1), which had a genome size of 5.84 megabytes (8). N. Azollae possessed a small G+C content of only 38.3%, and carried with it 4 rRNA clusters alongside 44 tRNA species, making up the full set of amino acids. Notable within the gene is that intact genes, as opposed to pseudogenes, compose 52% of the genome, which is the lowest of any cyanobacteria sequenced thus far, owing partly to the massive amount of pseudogenes that comprise the rest of the genome. In addition, N. Azollae has the some of the lowest intact CDS counts of any similar cyanobacteria sequenced (9).
Cell Structure, Metabolism and Life Cycle
Interesting features of cell structure; how it gains energy; what important molecules it produces.
Ecology and Pathogenesis
N. azollae, like most of the Nostoc genus, inhabits freshwater, tropical, temperate, and polar terrestrial systems, but are rarely found in marine environments (2).
Habitat; symbiosis; biogeochemical significance; contributions to environment.
If relevant, how does this organism cause disease? Human, animal, plant hosts? Virulence factors, as well as patient symptoms.
References
https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0011486
https://www.uniprot.org/proteomes/UP000001511
https://www.nobanis.org/globalassets/speciesinfo/a/azolla-filiculoides/azolla_filiculoides.pdf
https://sites.duke.edu/pryerlab/files/2019/02/Ariana_Azolla%E2%80%93Nostoc_2019.pdf
6. https://digitalcommons.wku.edu/cgi/viewcontent.cgi?article=1467&context=stu_hon_theses
https://www.sciencedirect.com/topics/agricultural-and-biological-sciences/nostoc
8. Handbook of Cyanobacteria
9. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0011486
Author
Page authored by John Weglarz, student of Prof. Jay Lennon at IndianaUniversity.