User:S4355920: Difference between revisions
| Line 30: | Line 30: | ||
==Cell structure and metabolism== | ==Cell structure and metabolism== | ||
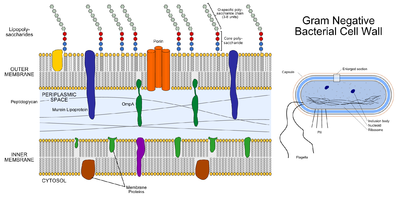
'''Cell Wall:''' P. gingivalis is an obligately aerobic, non-motile gram-negative bacterium <sup>[[#References|[2]]]</sup>. The cell wall is characterised by three distinct layers, including two membranous structures known as the inner membrane (IM) and the outer membrane (OM) <sup>[[#References|[12]]]</sup>. Connecting the two layers is a gel like structure known as the periplasm and a thin layer of peptidoglycan <sup>[[#References|[12]]]</sup>. The IM and OM possess a trilamellar structure composed of phospholipids <sup>[[#References|[12]]]</sup>. Distributed along the outer membrane are lipoproteins and lipopolysaccharides (LPS), which serve as an anchor for lipids <sup>[[#References|[12]]]</sup>. Chemically LPS is composed of three subunits, the O specific polysaccharide chain, the core and lipid A <sup>[[#References|[12]]]</sup>. | '''Cell Wall:''' P. gingivalis is an obligately aerobic, non-motile gram-negative bacterium <sup>[[#References|[2]]]</sup>. The cell wall is characterised by three distinct layers, including two membranous structures known as the inner membrane (IM) and the outer membrane (OM) <sup>[[#References|[12]]]</sup>. Connecting the two layers is a gel like structure known as the periplasm and a thin layer of peptidoglycan <sup>[[#References|[12]]]</sup>. The IM and OM possess a trilamellar structure composed of phospholipids <sup>[[#References|[12]]]</sup>. Distributed along the outer membrane are lipoproteins and lipopolysaccharides (LPS), which serve as an anchor for lipids <sup>[[#References|[12]]]</sup>. Chemically LPS is composed of three subunits, the O specific polysaccharide chain, the core and lipid A <sup>[[#References|[12]]]</sup>. | ||
[[File:Cell Wall GN.png|thumb|400 px| alt=Puzzle globe|Wikipedia logo|Gram negative cell wall structure]] | [[File:Cell Wall GN.png|thumb|400 px| alt=Puzzle globe|Wikipedia logo|Gram negative cell wall structure]] | ||
'''Fimbriae:''' Protruding the outer membrane of the cell wall, thin proteinaceous surface appendages aid and mediate bacterial attachment to the host <sup>[[#References|[8]]]</sup>. Approximately 25 μm long these structures have a robust ability to interact with salivary proteins, epithelial cells, extracellular matrix proteins and the fibroblasts of the host <sup>[[#References|[8]]]</sup>. Two distinct fimbriae types are displayed on the cell surfaces of the bacteria, known as FimA and Mfa protein <sup>[[#References|[8]]]</sup>. These surface structures are proposed to have a role in the progression of periodontal inflammatory reactions. Six genotypes of FimA structures exist (type I-V and Ib), ranging from 40.5 to 49kDa in size <sup>[[#References|[8]]]</sup>. Strain W83 is classed under type IV and are poorly fimbriated whereas strain ATCC 33277 are an abundantly fimbriated type I strain <sup>[[#References|[8]]]</sup>. The progression of chronic periodontitis is most closely associated with type II strains followed by type IV <sup>[[#References|[8]]]</sup>. | '''Fimbriae:''' Protruding the outer membrane of the cell wall, thin proteinaceous surface appendages aid and mediate bacterial attachment to the host <sup>[[#References|[8]]]</sup>. Approximately 25 μm long these structures have a robust ability to interact with salivary proteins, epithelial cells, extracellular matrix proteins and the fibroblasts of the host <sup>[[#References|[8]]]</sup>. Two distinct fimbriae types are displayed on the cell surfaces of the bacteria, known as FimA and Mfa protein <sup>[[#References|[8]]]</sup>. These surface structures are proposed to have a role in the progression of periodontal inflammatory reactions. Six genotypes of FimA structures exist (type I-V and Ib), ranging from 40.5 to 49kDa in size <sup>[[#References|[8]]]</sup>. Strain W83 is classed under type IV and are poorly fimbriated whereas strain ATCC 33277 are an abundantly fimbriated type I strain <sup>[[#References|[8]]]</sup>. The progression of chronic periodontitis is most closely associated with type II strains followed by type IV <sup>[[#References|[8]]]</sup>. | ||
'''Biofilm formation:''' The bacterium colonises the oral cavity by forming a complex biofilm known as plaque <sup>[[#References|[13]]]</sup>. They are recognised as secondary or late colonisers and require antecedent organisms to form the necessary environmental conditions for growth <sup>[[#References|[8]]]</sup>. Upon contact the bacterium must resist the plethora of host responses working against bacterial colonisation <sup>[[#References|[13]]]</sup>. Host factors are known to include mechanical shearing produced from the force of the tongue, saliva and gingival crevicular fluid flow <sup>[[#References|[13]]]</sup>. Successful colonisers must therefore possess a diverse repertoire of virulence factors to overcome host defences <sup>[[#References|[13]]]</sup>. | '''Biofilm formation:''' The bacterium colonises the oral cavity by forming a complex biofilm known as plaque <sup>[[#References|[13]]]</sup>. They are recognised as secondary or late colonisers and require antecedent organisms to form the necessary environmental conditions for growth <sup>[[#References|[8]]]</sup>. Upon contact the bacterium must resist the plethora of host responses working against bacterial colonisation <sup>[[#References|[13]]]</sup>. Host factors are known to include mechanical shearing produced from the force of the tongue, saliva and gingival crevicular fluid flow <sup>[[#References|[13]]]</sup>. Successful colonisers must therefore possess a diverse repertoire of virulence factors to overcome host defences <sup>[[#References|[13]]]</sup>. | ||
'''Motility:''' Non-motile <sup>[[#References|[3]]]</sup> | '''Motility:''' Non-motile <sup>[[#References|[3]]]</sup> | ||
'''Metabolic Functions:''' P. gingivalis is dependent on nitrogenous substrates for energy production (<sup>[[#References|[13]]]</sup>. Despite the nitrogenous compounds present in the oral cavity, the bacterium has a limited ability to ferment free amino acids <sup>[[#References|[13]]]</sup>. Aspartic acid and Asparagine are among the few, which can be metabolised to yield succinate. | '''Metabolic Functions:''' P. gingivalis is dependent on nitrogenous substrates for energy production (<sup>[[#References|[13]]]</sup>. Despite the nitrogenous compounds present in the oral cavity, the bacterium has a limited ability to ferment free amino acids <sup>[[#References|[13]]]</sup>. Aspartic acid and Asparagine are among the few, which can be metabolised to yield succinate. | ||
Revision as of 01:41, 15 September 2016
Name Bench ID Date MICR3004
Classification
Higher order taxa
Bacteria – Bacteroidetes – Bacteroidia – Bacteroidales – Porphyromonadaceae - Porphyromonas - P. gingivalis
Species
Species: Porphyromonas gingivalis
Strain: 2561 = ATCC 33277 = CCUG 25893 = CCUG 25928 = CIP 103683 = DSM 20709 = JCM 12257 = NCTC 11834
Description and significance
Porphyromonas gingivalis is an obligately aerobic, gram-negative bacterium belonging to the phylum Bacteroidetes [2]. Characterised by its rod shaped morphology, it is a non-spore bearing and non-motile bacterium most commonly inhabiting the oral cavity [2]. Recognised as an opportunistic pathogen, P. gingivalis is capable of living in commensal harmony with the host [3]. Termed as a pathobiont, the bacterium can cause episodes of diseases when a change in the ecological balance of the periodontal microenvironment transpires [4] [5]. Although the bacterium is capable of existing as a commensal organism, certain strains are known to be more virulent and pathogenic than others [3]. Virulent strains have found to include, W83, W50, ATCC 49417 and A7A1 [6] [7]. Avirulent strains include ATCC 381, 33277 and 23A4 [6] [7]. In vitro studies of the bacterium have found cells cultured in broth with a size range from 0.5 by 1 to 2 μm [2]. Cells grown on a solid media showed coccobacilli or very short rod structures (2). On blood agar plates, the bacterium forms black-pigmented colonies, predominately smooth, shiny and convex with a diameter between 1 to 2 mm [2] [8],.
Typically found in the oral cavity of individuals, P. gingivalis has been implicated with periodontal diseases, most commonly associated with chronic periodontitis [3]. A report from the Centres for Disease Control and Prevention (CDC) recorded 47. 2 % of adults in the United Stated aged 30 years or older have experienced some form of periodontal disease [9]. In light of this information, recent studies have also reported that P. gingivalis is associated with systematic diseases, including cardiovascular diseases, rheumatoid arthritis and decreased kidney function [10]. Studies underlying the molecular mechanisms behind the bacterial pathogenesis are key to design effective treatments. Consequently reducing the potential development of inflammatory diseases that arise as a secondary consequence of periodontitis.
Genome structure
The genome of strain W88 is comprised of a circular chromosome made up of 2 343 479 bp’s [11]. On average the guanine and cytosine content make up approximately 48.3 % of the genome [11].. The circular chromosome encodes 1909 protein genes 65 RNA genes [11]. 4 ribosomal operons (rrn, 5S rRNA-23S rRNA-tRNAAla-tRNAIle-16S rRNA) including 53 tRNA genes showing specificity for all 20 amino acids have been documented [11]. Interestingly the number of rrn operons and tRNA genes in strain W83 were identical to those of an avirulent strain counterpart ATCC 33277 [7] [11]. Nonetheless the extensive rearrangement between the two strains through the introduction of mobile elements inevitably altered the virulence of the bacterium [11].
The genome of W83 is composed predominately (85%) of ORF [11]. Encoding a total of 1,990 ORF, 1075 presented detectable biological roles [11]. Of the remaining ORF, 184 were categorised as a conserved hypothetical protein or conserved domain protein, 208 had to known function, and 523 encoded hypothetical proteins [11].
Cell structure and metabolism
Cell Wall: P. gingivalis is an obligately aerobic, non-motile gram-negative bacterium [2]. The cell wall is characterised by three distinct layers, including two membranous structures known as the inner membrane (IM) and the outer membrane (OM) [12]. Connecting the two layers is a gel like structure known as the periplasm and a thin layer of peptidoglycan [12]. The IM and OM possess a trilamellar structure composed of phospholipids [12]. Distributed along the outer membrane are lipoproteins and lipopolysaccharides (LPS), which serve as an anchor for lipids [12]. Chemically LPS is composed of three subunits, the O specific polysaccharide chain, the core and lipid A [12].
Fimbriae: Protruding the outer membrane of the cell wall, thin proteinaceous surface appendages aid and mediate bacterial attachment to the host [8]. Approximately 25 μm long these structures have a robust ability to interact with salivary proteins, epithelial cells, extracellular matrix proteins and the fibroblasts of the host [8]. Two distinct fimbriae types are displayed on the cell surfaces of the bacteria, known as FimA and Mfa protein [8]. These surface structures are proposed to have a role in the progression of periodontal inflammatory reactions. Six genotypes of FimA structures exist (type I-V and Ib), ranging from 40.5 to 49kDa in size [8]. Strain W83 is classed under type IV and are poorly fimbriated whereas strain ATCC 33277 are an abundantly fimbriated type I strain [8]. The progression of chronic periodontitis is most closely associated with type II strains followed by type IV [8].
Biofilm formation: The bacterium colonises the oral cavity by forming a complex biofilm known as plaque [13]. They are recognised as secondary or late colonisers and require antecedent organisms to form the necessary environmental conditions for growth [8]. Upon contact the bacterium must resist the plethora of host responses working against bacterial colonisation [13]. Host factors are known to include mechanical shearing produced from the force of the tongue, saliva and gingival crevicular fluid flow [13]. Successful colonisers must therefore possess a diverse repertoire of virulence factors to overcome host defences [13].
Motility: Non-motile [3]
Metabolic Functions: P. gingivalis is dependent on nitrogenous substrates for energy production ([13]. Despite the nitrogenous compounds present in the oral cavity, the bacterium has a limited ability to ferment free amino acids [13]. Aspartic acid and Asparagine are among the few, which can be metabolised to yield succinate.
Ecology
Pathology
Application to biotechnology
Bioengineering, biotechnologically relevant enzyme/compound production, drug targets,…
Current research
Summarise some of the most recent discoveries regarding this species.
References
8. [1]
9. [2]
- Chapter Book
This page is written by Amy Pham for the MICR3004 course, Semester 2, 2016