User:S4355920
Name Bench ID Date MICR3004
Classification
Higher order taxa
Bacteria – Bacteroidetes – Bacteroidia – Bacteroidales – Porphyromonadaceae - Porphyromonas - P. gingivalis
Species
Species: Porphyromonas gingivalis
Strain: 2561 = ATCC 33277 = CCUG 25893 = CCUG 25928 = CIP 103683 = DSM 20709 = JCM 12257 = NCTC 11834
Description and significance
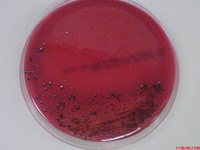
Porphyromonas gingivalis is an obligately aerobic, gram-negative bacterium belonging to the phylum Bacteroidetes [2]. Characterised by its rod shaped morphology, it is a non-spore bearing and non-motile bacterium most commonly inhabiting the oral cavity [2]. Recognised as an opportunistic pathogen, P. gingivalis is capable of living in commensal harmony with the host [3]. Termed as a pathobiont, the bacterium can cause episodes of diseases when a change in the ecological balance of the periodontal microenvironment transpires [4] [5]. Although the bacterium is capable of existing as a commensal organism, certain strains are known to be more virulent and pathogenic than others [3]. Virulent strains have found to include, W83, W50, ATCC 49417 and A7A1 [6] [7]. Avirulent strains include ATCC 381, 33277 and 23A4 [6] [7]. In vitro studies of the bacterium have found cells cultured in broth with a size range from 0.5 by 1 to 2 μm [2]. Cells grown on a solid media showed coccobacilli or very short rod structures (2). On blood agar plates, the bacterium forms black-pigmented colonies, predominately smooth, shiny and convex with a diameter between 1 to 2 mm [2] [8],.
Typically found in the oral cavity of individuals, P. gingivalis has been implicated with periodontal diseases, most commonly associated with chronic periodontitis [3]. A report from the Centres for Disease Control and Prevention (CDC) recorded 47. 2 % of adults in the United Stated aged 30 years or older have experienced some form of periodontal disease [9]. In light of this information, recent studies have also reported that P. gingivalis is associated with systematic diseases, including cardiovascular diseases, rheumatoid arthritis and decreased kidney function [10]. Studies underlying the molecular mechanisms behind the bacterial pathogenesis are key to design effective treatments. Consequently reducing the potential development of inflammatory diseases that arise as a secondary consequence of periodontitis.
Genome structure
The genome of strain W88 is comprised of a circular chromosome made up of 2 343 479 bp’s [11]. On average the guanine and cytosine content make up approximately 48.3 % of the genome [11].. The circular chromosome encodes 1909 protein genes 65 RNA genes [11]. 4 ribosomal operons (rrn, 5S rRNA-23S rRNA-tRNAAla-tRNAIle-16S rRNA) including 53 tRNA genes showing specificity for all 20 amino acids have been documented [11]. Interestingly the number of rrn operons and tRNA genes in strain W83 were identical to those of an avirulent strain counterpart ATCC 33277 [7] [11]. Nonetheless the extensive rearrangement between the two strains through the introduction of mobile elements inevitably altered the virulence of the bacterium [11].
The genome of W83 is composed predominately (85%) of ORF [11]. Encoding a total of 1,990 ORF, 1075 presented detectable biological roles [11]. Of the remaining ORF, 184 were categorised as a conserved hypothetical protein or conserved domain protein, 208 had to known function, and 523 encoded hypothetical proteins [11].
Cell structure and metabolism
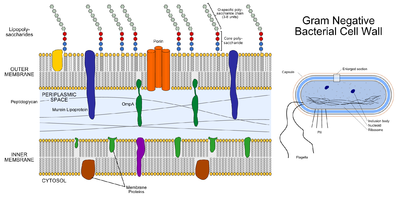
Cell Wall: P. gingivalis is an obligately aerobic, non-motile gram-negative bacterium [2]. The cell wall is characterised by three distinct layers, including two membranous structures known as the inner membrane (IM) and the outer membrane (OM) [12]. Connecting the two layers is a gel like structure known as the periplasm and a thin layer of peptidoglycan [12]. The IM and OM possess a trilamellar structure composed of phospholipids [12]. Distributed along the outer membrane are lipoproteins and lipopolysaccharides (LPS), which serve as an anchor for lipids [12]. Chemically LPS is composed of three subunits, the O specific polysaccharide chain, the core and lipid A [12].
Fimbriae: Protruding the outer membrane of the cell wall, thin proteinaceous surface appendages aid and mediate bacterial attachment to the host [8]. Approximately 25 μm long these structures have a robust ability to interact with salivary proteins, epithelial cells, extracellular matrix proteins and the fibroblasts of the host [8]. Two distinct fimbriae types are displayed on the cell surfaces of the bacteria, known as FimA and Mfa protein [8]. These surface structures are proposed to have a role in the progression of periodontal inflammatory reactions. Six genotypes of FimA structures exist (type I-V and Ib), ranging from 40.5 to 49kDa in size [8]. Strain W83 is classed under type IV and are poorly fimbriated whereas strain ATCC 33277 are an abundantly fimbriated type I strain [8]. The progression of chronic periodontitis is most closely associated with type II strains followed by type IV [8].
Biofilm formation: The bacterium colonises the oral cavity by forming a complex biofilm known as plaque [13]. They are recognised as secondary or late colonisers and require antecedent organisms to form the necessary environmental conditions for growth [8]. Upon contact the bacterium must resist the plethora of host responses working against bacterial colonisation [13]. Host factors are known to include mechanical shearing produced from the force of the tongue, saliva and gingival crevicular fluid flow [13]. Successful colonisers must therefore possess a diverse repertoire of virulence factors to overcome host defences [13].
Motility: Non-motile [3]
Metabolic Functions: P. gingivalis is dependent on nitrogenous substrates for energy production ([13]. Despite the nitrogenous compounds present in the oral cavity, the bacterium has a limited ability to ferment free amino acids [13]. Aspartic acid and Asparagine are among the few, which can be metabolised to yield succinate.
Ecology
Anaerobe: In the literature, P. gingivalis was previously regarded as a strictly anaerobic bacterium [14]. However current understanding of its genome composition indicates the presence of an oxygen metabolism pathway [14]. Although the bacterium grows optimally in aerobic conditions, the route towards the periodontal pockets involves exposure to oxygen [14]. Oxidative stress mechanisms are most likely employed to cope with the brief exposure to oxygen. Analysis of genome composition, revealed the presence of aerobic respiration genes (cydA and cydB), encoding the haem protein cytochrome bd oxidase [14].
Habitat: P. gingivalis is a natural inhabitant of the oral microbiome and is typically found residing in the subgingival sulcus of the oral cavity [5] [15]. The bacterium can also be found along the cheek, gingiva and tongue [3]. Although colonisation of the gingival crevice and periodontal pocket are typically associated with infection, colonisation of remote surfaces can also lead to disease [3]. Recognised as a secondary or late coloniser, attachment to antecedent organisms is typically evident [8]. Primary organisms provide a foundation for attachment and form the necessary environmental conditions for growth. Such as supplying growth substrates and decreasing oxygen tension [3] [8] Attachment to early plaque organisms includes species of oral streptococci (Streptococcus gordonii, S. sanguis, S. oralis, S. mitis, and S. crista) and Actinomyces naeslundii [3] [16] [17] [18] [19].
Microbe/Host Interaction:
Pathology
Application to biotechnology
Bioengineering, biotechnologically relevant enzyme/compound production, drug targets,…
Current research
Summarise some of the most recent discoveries regarding this species.
References
8. [1]
9. [2]
- Chapter Book
This page is written by Amy Pham for the MICR3004 course, Semester 2, 2016