Ostreococcus lucimarinus
Introduction
By Sarah Dendy
Ostreococcus are the smallest known eukaryotes1. They are single-celled but contain membrane-bound nuclei and a single chloroplast. They are approximately 1 μm in diameter, and are considered picophytoplankton.
Genomics
Ostreococcus lucimarinus, and a related species, Ostreococcus tauri, have been fully genome-sequenced. The genome of O. lucimarinus is 13.2 million base pairs, distributed on 21 chromosomes1, and is estimated to contain 7,651 genes1. High levels of similarity between O. lucimarinus and O. tauri provide insights to potential evolutionary mechanisms of the sister taxa. O. tauri has a genome of 12.6 million base pairs, which code an estimated 7,892 genes. Of these, 6,753 are orthologous to those of O lucimarinus. O tauri has 20 chromosomes, of which eighteen correspond very closely to those of O. lucimarinus. The other three O. lucimarinus chromosomes do not resemble those of its sister species, nor do they closely resemble anything else in the Ostreococcus genome. The origin of these chromosomes is somewhat mysterious. Chromosome 18, one such mysterious chromosome in O. lucimarinus, is the smallest chromosome of the organism. Its nucleotide sequence aligns with that of O. tauri chromosome 19, with about 5% of nucleotide sequence corresponding between the two species. By contrast, in other chromosomes, these species had 80-90% sequence homology. Chromosome 18 contains an estimated 83 genes, many of which code for proteins that most closely resemble those of bacteria. One enzyme, OSTLU28425, closely resembles the UDP-N-acetylglucosamine 2-epimerase of Microscilla marina, a marine bacterium. Palenik et al (2007) interpret this to indicate recent horizontal gene transfer between the planktonic bacterium and the miniscule planktonic eukaryote. They go on to propose that horizontal gene transfer between planktonic organisms, particularly in genes for sugar-processing or packaging pathways which could alter cell-surface composition and potentially hinder grazers or otherwise change microecological dynamics, will become a recurring theme in plankton genomics.
Micronutrients
The way in which Ostreococcus processes micronutrients, including metals, distinguishes it from other plankton. O. lucimarinus creates a large number of selenocysteine-containing enzymes (selenoproteins), which are enzymes whose catalytic activity is heightened by substituting a cysteine in an active site to a selenocysteine.[1] Theoretically, this substitution allows cells to have heightened activity from a single enzyme, and therefore need to manufacture fewer enzymes to achieve the same physiological effects. In highly-expressed enzymes, this allows a cell to save molecular resources like nitrogen, because the manufacture of fewer proteins translates to fewer nitrogen atoms being associated with protein backbone. This allows cells to more efficiently use amino acids, which may be valuable at the extremes of livable conditions. However, selenoprotein abundance is limited by availability of selenium, as well as by the evolution of complex recognition systems in coding DNA, which must affix a selenocysteine where typical DNA machinery would recognize a stop codon (TGA). Though iron is a limiting nutrient in many ecosystems, including in marine planktonic ecosystems, Ostreococcus has no described system of iron uptake analogous to those of related organisms, like diatoms. Ostreococcus has no ferric reductase, multicopper oxidase, or ferric permease, all of which are common elements of eukaryotic iron uptake systems (though O. tauri may have a multicopper oxidase, which is not found in any other lineage of the genus). Predicted adaptations to low iron levels are not found in O. lucimarinus. Several iron atoms are required for molecules critical to photosynthesis, the organism’s main means of survival. Also of note, O lucimarinus lacks systems for responding to high levels of copper toxicity through a phytochelatin synthase. This organism presumably must have novel ways of responding to low iron levels, or of responding to copper toxicity, but they are not currently known or described. Furthermore, Ostreococcus requires some of its micronutrients, like vitamin B12, from the environment, because it lacks to genetic pathways to endogenously synthesize this nutrient, but still depends upon it for other physiological functions.
Evolution of the Genus
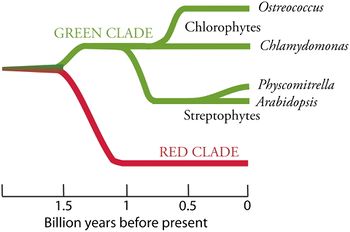
Ostreococcus has several adaptations specific to allowing its incredibly small size. Comparisons to Chlamydomonas, a closely related genus of photosynthetic diatoms, reveals a large number of genes which are common and well-characterized through the plant kingdom. Some of these well-known genes are nonetheless absent in Ostreococcus, but because Chlamydomonas demonstrates that they already existed in an ancestral phytoplankton phase, a fair assumption is that they have been lost in Ostreococcus [2]. Fascinatingly, one gene which is found in the organelles of plants and algae is actually found in the nuclear genome of Ostreococcus. This gene, CcsA, codes a hydrophobic protein significant to the handling of heme groups in system II cytochrome biogenesis. This is the first described example of presumed gene transfer from the organelle of a cell to the cell nucleus. The implications of this observation are enormous, because organelle genomes are used in countless cell lineages for developing phylogenetic and evolutionary relationships. If horizontal gene transfer, as documented here, occurs between organelles and cell nuclei, many existing taxonomic analyses could be subject to revision with the inclusion of nuclear data. However, this may be a relatively rare occurrence, and perhaps only possible in Ostreococcus because the extreme cell size puts selective pressure on novel gene dynamics and behaviors. Furthermore, probably because pressure toward small cell size has pressured these organisms to reduce their amount of genetic material, Ostreococcus has developed a number of fusion proteins. Around 348 fusion proteins were identified from O. lucimarinus, of which 49 are known to combine multiple enzymatic functions, and the remainder may be either true fusions or combinations of anticipated genes, without additive enzymatic functions. Interestingly, some fusions occur between proteins originating from different biochemical pathways. Also of interest, despite the pressure to reduce genome size through such apparently drastic means as fusing functional proteins, O. lucimarinus still has introns and noncoding sequences in its nuclear DNA. Though several other eukaryotic characteristics, such as broadly-shared chromatin proteins, have been lost in Ostreococcus, the eukaryotic capacity for post-transcriptional splicing remains in this lineage. GapB2 is a gene found in Ostreococcus with significant implications for the evolution of the genus. GapB is an abbreviation for glyceraldehyde-3-phosphate dehydrogenase, which is a cell-signaling component for down-regulating the Calvin cycle in land plants during the night. GapB originates from the duplication of another glyceraldehyde-3-phosphate dehydrogenase gene, known as GapA, with a modification on its C-terminal end from a gene with related function, known as CP12. However, Ostreococcus provides evidence against the duplication-hypothesis for the origin of GapB, because it contains a copy of this gene, as does Mesostigma viride, a phylogenetically separate green algae. If the duplication had arisen once, GapB should appear in only one of these lineages, not both. The fact of its presence may be another example of lateral gene transfer, which is the best existing explanation for several genetic components of the Ostreococccus genus. More data on the rate and character of lateral gene transfer in these organisms would be invaluable to an understanding of their evolution, as well as life histories. Significantly, lateral gene transfer of UDP-N-acetylglucosamine 2-epimerase to Ostreococcus is anticipated from a bacterium, whereas transfer of glyceraldehyde-3-phosphate dehydrogenase B is from an algae, another eukaryote. Also interestingly, Ostreococcus lacks the CP12 gene, which functions in a role similar to that of GapB. Ostreococcus has a genome reduced in size in many areas; possibly, the deletion of CP12 is a continuation on this theme, and represents an instance in which lateral gene transfer allows the recipient organism to capitalize on its new gene repertoire to streamline its genome overall. Yet another protein group valuable for evolutionary study is the polyketide synthase enzymes3, relevant to lipid metabolism. Polyketide synthases are found in Ostreococcus, as well as in the related genus Chlamydomonas. This is extremely unusual, because these enzymes typically occur in fungi and bacteria. Though intuitively this should represent an instance of lateral gene transfer, in fact the polyketide synthases of protists, including Ostreococcus, differ from those of all potential donor lineages. Therefore, it is unlikely that Ostreococcus polyketide synthase originated from a marine bacterium. Whether this enzyme group evolved spontaneously in Ostreococcus and Chlamydomonas, versus in other organisms, or whether it evolved once and was lost in a large number of eukaryotes, is difficult to ascertain from existing data, and not yet known.[3]
Introns
Because Ostreococcus is a genus of such small eukaryotes, they have an extremely condensed genome. Their density of genes, or ratio of coding sequence to noncoding or ‘junk’ DNA, is exceptionally high among eukaryotes. They conserve intracellular space in part by reducing the amount of nuclear DNA they must contain. Furthermore, the small genome size is evolutionarily derived, that is, in the ancestors of Ostreococcus genomes and cells were both larger in size, and selective pressure caused individuals to become reduced in size. Interestingly, this reduction does not apply to as great an extent to intragenic noncoding DNA, or introns. The intron density in Ostreococcus DNA is comparably high to that in other eukaryotes, including species with much larger genomes and much larger cells for containing their increased genetic code. A simple explanation for directional selection reducing size would suggest the elimination of introns at a comparable rate to the decrease in noncoding, intergenic DNA. The fact that this is not observed suggests a more complex role of introns. Introns may convey some selective advantage to genes containing them, and Ostreococcus provides an ideal study system for this dynamic. Furthermore, the localization of introns in the Ostreococcus genome is of interest. Introns are not equally likely to appear in all genes of the genome, but are biased toward genes which are expressed at a higher level, and away from genes with a low level of expression. The more frequently an Ostreococcus cell manufactures cellular machinery from a genetic sequence, the more likely it is that that genetic sequence contains untranslated introns. Thus, the Ostreococcus genome reflects a double bias, in that nucleic acid content is more likely to be deleted from intergenic regions than noncoding intragenic regions, and that noncoding nucleic acids are more likely to occur in genes which are frequently expressed than in genes that are not.[4] Just as the bias in sequence deletions indicates a selective advantage to introns, the position of Ostreococcus introns may reflect the character of the selective advantage. Existing Ostreococcus data suggests that introns convey greater advantage when the genes with which they are associated are frequently transcribed. The mechanism for this difference remains unclear; possible approaches to its study might examine the role of introns in this taxon in DNA transcription or translation, or in DNA maintenance, resilience to mutation, storage in small spaces, or other significant affected characteristics.
Poly-unsaturated Fat Chain Synthesis
Polyunsaturated fat chains (PUFAs) are a component of the human diet which is beneficial to human health. Ostreococcus lucimarinus is relevant to the study of these molecules because it can provide cDNA (coding DNA, or DNA sequence lacking noncoding introns) for several of the enzymes required for PUFA synthesis. PUFAs, including very long-chain PUFAs (VLCPUFAs) are synthesized endogenously in humans at only very low levels, so their supplementation from diet is extremely significant to human nutrition. Description of the VLCPUFA synthesis pathway allows for its optimization in crop plants or other nutrient sources in human diet. However, research on this synthesis pathway is challenging in most eukaryotes, because eukaryote genes contain a complex of introns and exons which obfuscate the true sequence, and therefore protein structure and function, of genes relevant to the synthesis pathway. Ostreococcus allows description of some of these synthesis pathways in a relatively simplified, and easier to manipulate, system than the crop plants themselves. One such cDNA isolated from Ostreococcus is Δ4-fatty acid desaturase, an enzyme catalyzing desaturation of substrates, especially VLCPUFAs.[5]
Algae-specific Viruses
Many outstanding questions accompany the modern understanding of the diversity of eukaryotic viruses. One extremely poorly-understood system is the study of viral diversity in seawater. Viruses have the potential to be enormously diverse, and a marine system represents an excess of space for the proliferation and differentiation of viruses. However, there are no boundaries to viral dispersal in the ocean, and it is unclear to what extent global geography hinders viral homogeny, and to what extent viruses from one marine site are isolated from those in another. Planktonic organisms like Ostreococcus provide an interesting study system for hosts of marine viruses. Because planktonic species are widespread and common, they provide a substrate in which viral communities develop and specialize. Methodologies for isolating viral strains include lysis plaque assays against a lawn of putative host cell, like Ostreococcus lucimarinus. Some viral studies include comparisons within the Ostreococcus genus with related species like Ostreococcus tauri. O. lucimarinus is more widespread than O. tauri, and occupies a wider range of ecological niches. Consistent with this observation, O. tauri is documented to be affected by viruses only in water sampled near its natural habitat, whereas O. lucimarinus is vulnerable to viruses from a large range of seawater samples, including those much more distant from its natural growing conditions. Assays of one species against the other indicate different ecological dynamics reflected in viral phylogeny. The viruses infecting one species were not necessarily more diverse than the viruses affecting a single species but originating from different locations. Derelle et al (2015) described six strains of virus in two broad subgroups infecting O. lucimarinus, out of a probable assortment of a great number more than that worldwide.[6] The genomes of these viruses were highly similar overall, and viruses from each sampled seawater site were able to infect all studied O. lucimarinus. This result is consistent with the notion that viruses disperse freely in marine systems and the lack of geographic limitation results in a lack of barriers to viral efficacy in multiple systems. A virus in a single species in a given site may be genetically different from the virus of the same species at a different site, but it will have the ability to cross-infect the host from either location, implying a continuous ecology between sampling locations and throughout the ocean overall. Interesting areas for future viral study could include the evolutionary origins of the viruses, particularly whether viruses with one Ostreococcus host species switched at some point in their evolutionary history to the other. Other marine systems have also demonstrated other viral ecological dynamics such as seasonality. This has been suggested, but not explored or fully documented in the Ostreococcus lucimarinus viral suite.
FTO Obesity Gene and Homologies
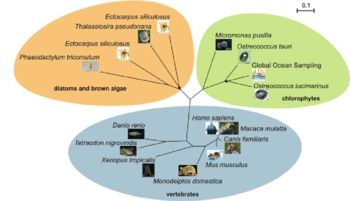
The FTO or fat mass and obesity associated gene is a risk factor gene for obesity and excessively high body mass index, which has an unexpected homology with genes in Ostreococcus lucimarinus. Monogenic obesity cases, that is, instances in which obesity is triggered by a single gene, are documented in a small portion of obesity sufferers, normally children. The vast majority of obesity patients have a genetic element predisposing them to the disease, but monogenic cases represent a tiny minority, and an overwhelming proportion of obese people have a polygenic predisposition, or a large group of genes together contributing to risk factors. This polygenic system is much more difficult to identify and address than a one-gene inheritance, but among implicated genes for obesity risk, FTO shows the strongest correlations with obesity diagnosis. Specifically, a forty-seven kilobase region in the first two introns of the FTO demonstrates high linkage disequilibrium and is associated strongly with obesity.[7] FTO is differentially expressed in different tissues. Studies in mice suggest that it causes phenotypic differences mostly when expressed in brain tissues, and may have age-dependent effects. The elements of the gene which are obesity-associated are in an intron, and therefore do not affect the structure of the protein as it is expressed. Introns have other roles in genes, including interacting with promoters or other transcription-regulatory effects. Because of their location near the beginning of the gene, in introns one and two, it seems likely that the FTO polymorphisms most associated with obesity affect a promoter or repressor of the FTO gene. The promoter IRX3, which is located on the same chromosome as FTO in mice but downstream by about half a megabase, has been documented interacting with the polymorphic, obesity-linked intron region of FTO in adult mouse brains[8]. The FTO gene promoter, by contrast, has interactions only very close to its own location on the chromosome, and does not interact with the obesity-linked intron region at all, except in developing, embryonic mouse brains. Both FTO and IRX3 are conserved among vertebrates. Their existence has not been documented in invertebrate animals, fungi, or higher plants, despite assays for homology in a range of completely-sequenced genomes. FTO research is somewhat stymied by the absence of FTO genes in a range of model organisms, like Drosophila melanogaster, Caenorhabditis elegans, Arabidopsis thaliana, and Chlamydomonas reinhardtii. Therefore, the discovery of homologies to FTO in several algal genomes comes as something of a surprise. Ostreococcus and the related Micromonas have a gene sequence homologous to that found in vertebrate FTO, as do several larger multicellular algae, like Phaeodactylum tricornutum and Ectocarpus siliculosus. FTO and FTO homologies are sorted loosely into three clades, one including vertebrate animals, one including brown algae, diatoms and multicellular algae like those listed here, and a third including picoplankton like Ostreococcus. Interestingly, no FTO homologs have yet been observed in freshwater algae; all documented algae with these homologies are marine species. The potential for Ostreococcus lucimarinus to serve as a model species for obesity research is highly uncertain at best. The exact role and mechanism of FTO, and the proteins it encodes, is poorly understood. FTO is differently expressed between different mammalian tissues, with significant expression in the brain and in glands associated with the endocrine system. Furthermore, significant regions of FTO for obesity-linkage analyses occur in introns, which suggest they may be significant to the interaction of other genes on the transcription of FTO; if these unknown other genes lack homology in Ostreococcus, it would limit the efficacy of the algae as a model for the study of this system. However, the unexpected appearance of gene homologs in this taxonomic group is exciting for potential research applications. Many algae are easily cultured in laboratory conditions, especially in comparison to animal models for FTO like mouse (Mus musculus) or zebrafish (Danio rerio). Algae in obesity research is a developing area.[9]
References
- ↑ Robbens, Steven, Petersen, Jörn, Brinkmann, Henner, Rouzé, Pierre, & van de Peer, Yves. (2006). Unique regulation of the Calvin Cycle in the ultrasmall green alga Ostreococcus. Journal of Molecular Evolution 64:601-604
- ↑ Palenik, Brian, Grimwood, Jane, Aerts, Andrea, Rouzé, Pierre, Salamov, Asaf, Putnam, Nicholas, Dupont, Chris, Jorgensen, Richard, Derelle, Evelyne, Rombauts, Stephane, Zhou, Kemin, Otillar, Robert, Merchant, Sabeeha S., Podell, Sheila, Gaasterland, Terry, Napoli, Carolyn, Gendler, Karla, Manuell, Andrea, Tai, Vera, Vallon, Olivier, Piganeau, Gwenael, Jancek, Séverine, Heijde, Marc, Jabbari, Kamel, Bowler, Chris, Lohr, Martin, Robbens, Steven, Werner, Gregory, Dubchak, Inna, Pazour, Gregory J., Ren, Qinghu, Paulsen, Ian, Delwiche, Chuck, Schmutz, Jeremy, Rokhsar, Daniel, van de Peer, Yves, Moreau, Hervé, & Grigoriev, Igor V. (2007) The tiny eukaryote Ostreococcus provides genomic insights into the paradox of plankton speciation. Proceedings of the National Academy of Sciences of the United States of America, 104 (18): 7705-7710
- ↑ John, Uwe, Beszteri, Bánk, Derelle, Evelyne, van de Peer, Yves, Read, Betsy, Moreau, Hervé, & Cembella, Allan (2008). Novel insights into evolution of protistan polyketide synthases through phylogenomic analysis. Protist, 159:21-30
- ↑ Lanier, William, Moustafa, Ahmed, Bhattacharya, Debashish, &Comeron, Josep M. (2008) EST analysis of Ostreococcus lucimarinus, the most compact eukaryotic genome, shows an excess of introns in highly expressed genes. PLoS ONE 3(5): e2171
- ↑ Ahmann, Katharina, Heilmann, Mareike, & Feussner, Ivo (2011) Identification of a Δ4-desaturase from the microalga Ostreococcus lucimarinus. European Journal of Lipid Science and Technology. 113: 832-840
- ↑ Derelle, Evelyne, Monier, Adam, Cooke, Richard, Worden, Alexandra Z., Grimsley, Nigel H., & Moreau, Hervé (2015). Diversity of viruses infecting the green microalga Ostreococcus lucimarinus, 89(11): 5812-5821
- ↑ Frayling, Timothy M., Timpson, Nicholas J., Weedon, Michael N., Zeggini, Eleftheria, Freathy, Rachel M., Lindgren, Cecilia M., Perry, John R. B., Elliott, Katherine S., Lango, Hana, Rayner, Nigel W., Shields, Beverley, Harries, Lorna W., Barrett, Jeffrey C., Ellard, Sian, Groves, Christopher J., Knight, Bridget, Patch, Ann-Marie, Ness, Andrew R., Ebrahim, Shah, Lawlor, Debbie A., Ring, Susan M., Ben-Shlomo, Yoav, Jarvelin, Marjo-Riitta, Sovio, Ulla, Bennett, Amanda J., Melzer, David, Ferrucci, Luigi, Loos, Ruth J. F., Barroso, Iñes, Wareham, Nicholas J., Karpe, Fredrik, Owen, Katharine R., Cardon, Lon R., Walker, Mark, Hitman, Graham A., Palmer, Colin N. A., Doney, Alex S. F., Morris, Andrew D., Davey Smith, George, The Wellcome Trust Case Control Consortium, Hattersley, Andrew T., & McCarty, Mark I. (2007) A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science, 316(5826):889-894
- ↑ Smemo, Scott, Tena, Juan J., Kim, Kyoung-Han, Gamazon, Eric R., Sakabe, Noboru J., Gómez-Marín, Carlos, Aneas, Ivy, Credidio, Flavia L., Sobreira, Débora R., Wasserman, Nora F., Lee, Ju Hee, Puviindran, Vijitha, Tam, Davis, Shen, Michael, Son, Joe Eun, Vakili, Niki Alizadeh, Sung, Hoon-Ki, Naranjo, Silvia, Acemel, Rafael D., Manzanares, Miguel, Nagy, Andras, Cox, Nancy J., Hui, Chi-Chung, Gomez-Skarmeta, Jose Luis, &Nóbrega, Marcelo A. (2014). Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature, 507(7492): 371-375
- ↑ Robbens, Steven, Rouzé, Pierre, Cock, J. Mark, Spring, Jürg, Worden, Alexandra Z., & van de Peer, Yves (2008). The FTO gene, implicated in human obesity, is found only in vertebrates and marine algae. Journal of Molecular Evolution 66: 80-84
Authored for BIOL 238 Microbiology, taught by Joan Slonczewski, 2018, Kenyon College.